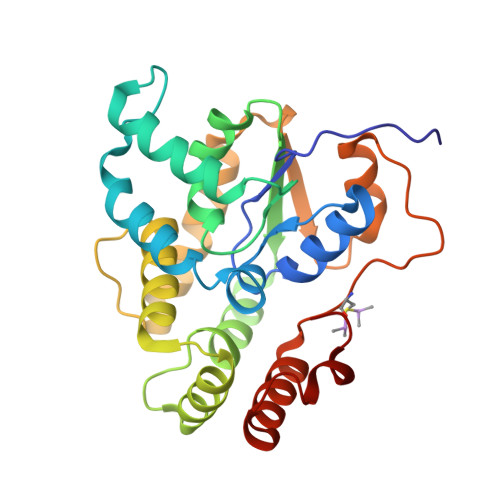
Oxamniquine derivatives overcome Praziquantel treatment limitations for Schistosomiasis.
Alwan, S.N., Taylor, A.B., Rhodes, J., Tidwell, M., McHardy, S.F., LoVerde, P.T.(2023) PLoS Pathog 19: e1011018-e1011018
- PubMed: 37428793 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1011018
- Primary Citation Related Structures:
8E5Q, 8E5R - PubMed Abstract:
Human schistosomiasis is a neglected tropical disease caused by Schistosoma mansoni, S. haematobium, and S. japonicum. Praziquantel (PZQ) is the method of choice for treatment. Due to constant selection pressure, there is an urgent need for new therapies for schistosomiasis. Previous treatment of S. mansoni included the use of oxamniquine (OXA), a drug that is activated by a schistosome sulfotransferase (SULT). Guided by data from X-ray crystallography and Schistosoma killing assays more than 350 OXA derivatives were designed, synthesized, and tested. We were able to identify CIDD-0150610 and CIDD-0150303 as potent derivatives in vitro that kill (100%) of all three Schistosoma species at a final concentration of 71.5 μM. We evaluated the efficacy of the best OXA derivates in an in vivo model after treatment with a single dose of 100 mg/kg by oral gavage. The highest rate of worm burden reduction was achieved by CIDD -150303 (81.8%) against S. mansoni, CIDD-0149830 (80.2%) against S. haematobium and CIDD-066790 (86.7%) against S. japonicum. We have also evaluated the ability of the derivatives to kill immature stages since PZQ does not kill immature schistosomes. CIDD-0150303 demonstrated (100%) killing for all life stages at a final concentration of 143 μM in vitro and effective reduction in worm burden in vivo against S. mansoni. To understand how OXA derivatives fit in the SULT binding pocket, X-ray crystal structures of CIDD-0150303 and CIDD-0150610 demonstrate that the SULT active site will accommodate further modifications to our most active compounds as we fine tune them to increase favorable pharmacokinetic properties. Treatment with a single dose of 100 mg/kg by oral gavage with co-dose of PZQ + CIDD-0150303 reduced the worm burden of PZQ resistant parasites in an animal model by 90.8%. Therefore, we conclude that CIDD-0150303, CIDD-0149830 and CIDD-066790 are novel drugs that overcome some of PZQ limitations, and CIDD-0150303 can be used with PZQ in combination therapy.
- Departments of Biochemistry and Structural Biology, University of Texas Health at San Antonio; San Antonio, Texas, Unites States of America.
Organizational Affiliation: