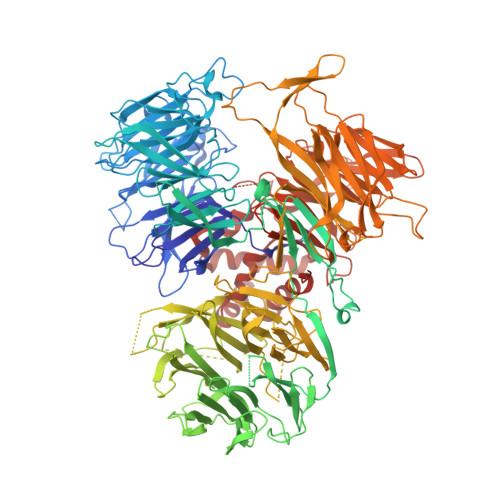
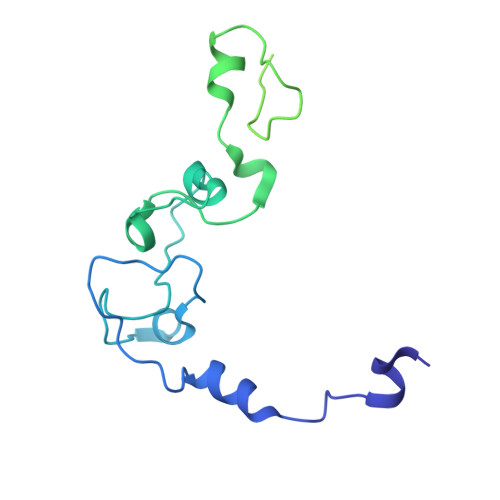
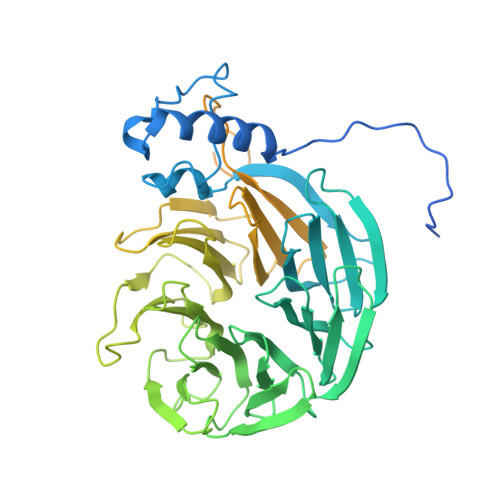
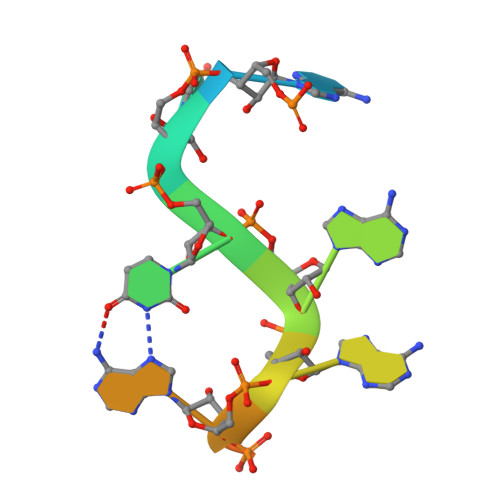
Molecular basis for the recognition of the AUUAAA polyadenylation signal by mPSF.
Gutierrez, P.A., Wei, J., Sun, Y., Tong, L.(2022) RNA 28: 1534-1541
- PubMed: 36130077 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.079322.122
- Primary Citation Related Structures:
8E3I, 8E3Q - PubMed Abstract:
The polyadenylation signal (PAS) is a key sequence element for 3'-end cleavage and polyadenylation of messenger RNA precursors (pre-mRNAs). This hexanucleotide motif is recognized by the mammalian polyadenylation specificity factor (mPSF), consisting of CPSF160, WDR33, CPSF30, and Fip1 subunits. Recent studies have revealed how the AAUAAA PAS, the most frequently observed PAS, is recognized by mPSF. We report here the structure of human mPSF in complex with the AUUAAA PAS, the second most frequently identified PAS. Conformational differences are observed for the A1 and U2 nucleotides in AUUAAA compared to the A1 and A2 nucleotides in AAUAAA, while the binding modes of the remaining 4 nt are essentially identical. The 5' phosphate of U2 moves by 2.6 Å and the U2 base is placed near the six-membered ring of A2 in AAUAAA, where it makes two hydrogen bonds with zinc finger 2 (ZF2) of CPSF30, which undergoes conformational changes as well. We also attempted to determine the binding modes of two rare PAS hexamers, AAGAAA and GAUAAA, but did not observe the RNA in the cryo-electron microscopy density. The residues in CPSF30 (ZF2 and ZF3) and WDR33 that recognize PAS are disordered in these two structures.
- Department of Biological Sciences, Columbia University, New York, New York 10027, USA.
Organizational Affiliation: