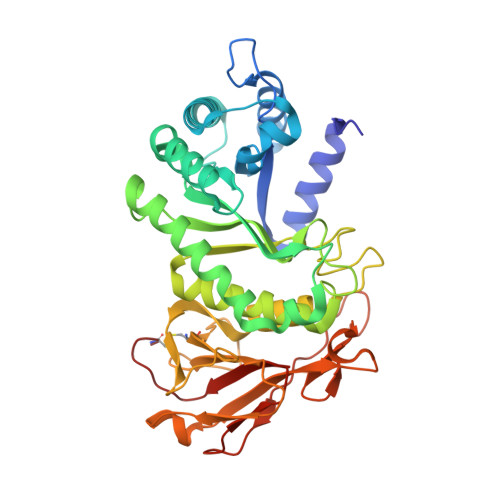
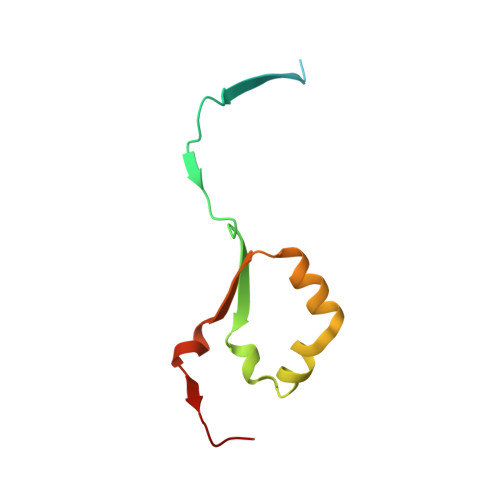
Complex Inhibitory Mechanism of Glycomimetics with Heparanase.
Whitefield, C., Vo, Y., Schwartz, B.D., Hepburn, C., Ahmed, F.H., Onagi, H., Banwell, M.G., Nelms, K., Malins, L.R., Jackson, C.J.(2023) Biochemistry 62: 2202-2215
- PubMed: 37368361 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.3c00038
- Primary Citation Related Structures:
8E07, 8E08 - PubMed Abstract:
Heparanase (HPSE) is the only mammalian endo -β-glucuronidase known to catalyze the degradation of heparan sulfate. Dysfunction of HPSE activity has been linked to several disease states, resulting in HPSE becoming the target of numerous therapeutic programs, yet no drug has passed clinical trials to date. Pentosan polysulfate sodium (PPS) is a heterogeneous, FDA-approved drug for the treatment of interstitial cystitis and a known HPSE inhibitor. However, due to its heterogeneity, characterization of its mechanism of HPSE inhibition is challenging. Here, we show that inhibition of HPSE by PPS is complex, involving multiple overlapping binding events, each influenced by factors such as oligosaccharide length and inhibitor-induced changes in the protein secondary structure. The present work advances our molecular understanding of the inhibition of HPSE and will aid in the development of therapeutics for the treatment of a broad range of pathologies associated with enzyme dysfunction, including cancer, inflammatory disease, and viral infections.
- Research School of Chemistry, Australian National University, Canberra, Australian Capital Territory 2601, Australia.
Organizational Affiliation: