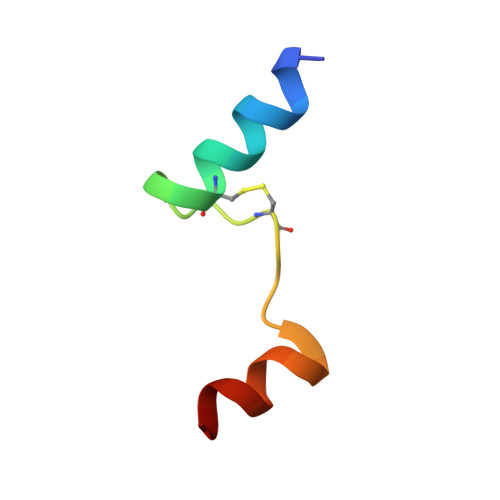
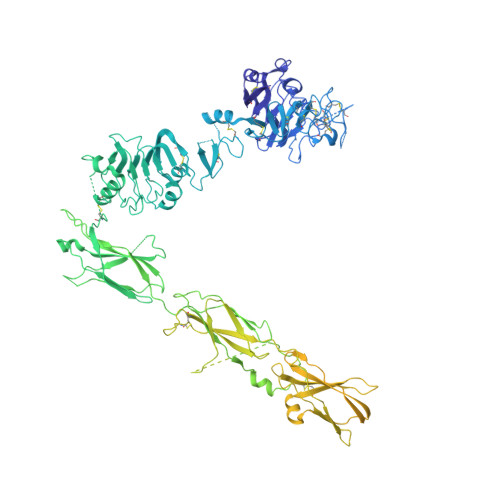
Activation of the insulin receptor by an insulin mimetic peptide.
Park, J., Li, J., Mayer, J.P., Ball, K.A., Wu, J., Hall, C., Accili, D., Stowell, M.H.B., Bai, X.C., Choi, E.(2022) Nat Commun 13: 5594-5594
- PubMed: 36151101 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-33274-0
- Primary Citation Related Structures:
8DTL, 8DTM - PubMed Abstract:
Insulin receptor (IR) signaling defects cause a variety of metabolic diseases including diabetes. Moreover, inherited mutations of the IR cause severe insulin resistance, leading to early morbidity and mortality with limited therapeutic options. A previously reported selective IR agonist without sequence homology to insulin, S597, activates IR and mimics insulin's action on glycemic control. To elucidate the mechanism of IR activation by S597, we determine cryo-EM structures of the mouse IR/S597 complex. Unlike the compact T-shaped active IR resulting from the binding of four insulins to two distinct sites, two S597 molecules induce and stabilize an extended T-shaped IR through the simultaneous binding to both the L1 domain of one protomer and the FnIII-1 domain of another. Importantly, S597 fully activates IR mutants that disrupt insulin binding or destabilize the insulin-induced compact T-shape, thus eliciting insulin-like signaling. S597 also selectively activates IR signaling among different tissues and triggers IR endocytosis in the liver. Overall, our structural and functional studies guide future efforts to develop insulin mimetics targeting insulin resistance caused by defects in insulin binding and stabilization of insulin-activated state of IR, demonstrating the potential of structure-based drug design for insulin-resistant diseases.
- Department of Pathology and Cell Biology, Vagelos College of Physicians and Surgeons, Columbia University, New York, NY, 10032, USA.
Organizational Affiliation: