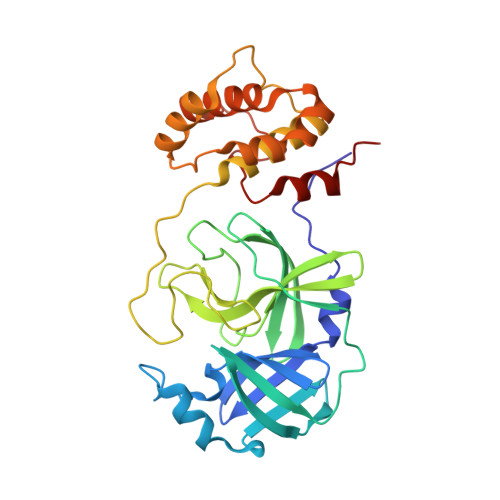
X-ray crystallographic characterization of the SARS-CoV-2 main protease polyprotein cleavage sites essential for viral processing and maturation.
Lee, J., Kenward, C., Worrall, L.J., Vuckovic, M., Gentile, F., Ton, A.T., Ng, M., Cherkasov, A., Strynadka, N.C.J., Paetzel, M.(2022) Nat Commun 13: 5196-5196
- PubMed: 36057636 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-32854-4
- Primary Citation Related Structures:
8DRR, 8DRS, 8DRT, 8DRU, 8DRV, 8DRW, 8DRX, 8DRY, 8DRZ, 8DS0, 8DS1, 8DS2 - PubMed Abstract:
Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2), the pathogen that causes COVID-19, produces polyproteins 1a and 1ab that contain, respectively, 11 or 16 non-structural proteins (nsp). Nsp5 is the main protease (M pro ) responsible for cleavage at eleven positions along these polyproteins, including at its own N- and C-terminal boundaries, representing essential processing events for viral assembly and maturation. Using C-terminally substituted M pro chimeras, we have determined X-ray crystallographic structures of M pro in complex with 10 of its 11 viral cleavage sites, bound at full occupancy intermolecularly in trans, within the active site of either the native enzyme and/or a catalytic mutant (C145A). Capture of both acyl-enzyme intermediate and product-like complex forms of a P2(Leu) substrate in the native active site provides direct comparative characterization of these mechanistic steps as well as further informs the basis for enhanced product release of M pro 's own unique C-terminal P2(Phe) cleavage site to prevent autoinhibition. We characterize the underlying noncovalent interactions governing binding and specificity for this diverse set of substrates, showing remarkable plasticity for subsites beyond the anchoring P1(Gln)-P2(Leu/Val/Phe), representing together a near complete analysis of a multiprocessing viral protease. Collectively, these crystallographic snapshots provide valuable mechanistic and structural insights for antiviral therapeutic development.
- Department of Biochemistry and Molecular Biology and Centre for Blood Research, The University of British Columbia, Vancouver, BC, V6T 1Z3, Canada.
Organizational Affiliation: