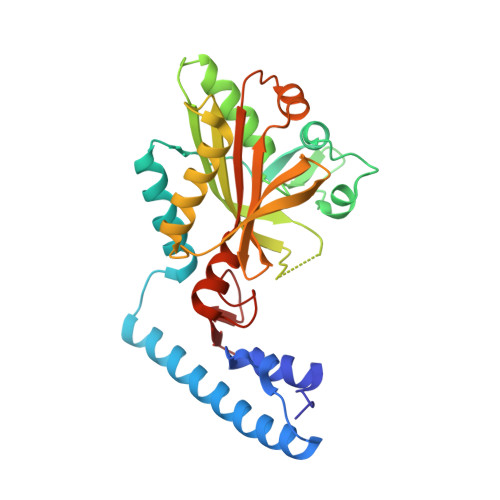
Structures of Methanomethylophilus alvus Pyrrolysine tRNA-Synthetases Support the Need for De Novo Selections When Altering the Substrate Specificity.
Gottfried-Lee, I., Perona, J.J., Karplus, P.A., Mehl, R.A., Cooley, R.B.(2022) ACS Chem Biol 17: 3470-3477
- PubMed: 36395426 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.2c00640
- Primary Citation Related Structures:
8DQG, 8DQH, 8DQI, 8DQJ - PubMed Abstract:
A recently developed genetic code expansion (GCE) platform based on the pyrrolysine amino-acyl tRNA synthetase (PylRS)/tRNA Pyl pair from Methanomethylophilus alvus (Ma) has improved solubility and lower susceptibility to proteolysis compared with the homologous and commonly used Methanosarcina barkeri (Mb) and M. mazei (Mm) PylRS GCE platforms. We recently created two new Ma PylRS variants for the incorporation of the fluorescent amino acid, acridonyl-alanine (Acd), into proteins at amber codons: one based on "transplanting" active site mutations from an established high-efficiency Mb PylRS and one that was de novo selected from a library of mutants. Here, we present the crystal structures of these two Ma PylRS variants with Acd/ATP bound to understand why the "active site transplant" variant (Acd-AST) displayed 6-fold worse Acd incorporation efficiency than the de novo selected PylRS (called Acd-RS1). The structures reveal that the Acd-AST binding pocket is too small and binds the three-ring aromatic Acd in a distorted conformation, whereas the more spacious Acd-RS1 active site binds Acd in a relaxed, planar conformation stabilized by a network of solvent-mediated hydrogen bonds. The poor performance of the AST enzyme is ascribed to a shift in the Ma PylRS β-sheet framework relative to that of the Mb enzyme. This illustrates a general reason why "active site transplantation" may not succeed in creating efficient Ma PylRSs for other noncanonical amino acids. This work also provides structural details that will help guide the development of future Ma PylRS/tRNA Pyl GCE systems via de novo selection or directed evolution methods.
- Department of Biochemistry and Biophysics, 2011 Agricultural and Life Sciences, Oregon State University, Corvallis, Oregon 97331, United States.
Organizational Affiliation: