Characterization of synthetic antigen binding fragments targeting Toc75 for the isolation of TOC in A. thaliana and P. sativum.
Srinivasan, K., Erramilli, S.K., Chakravarthy, S., Gonzalez, A., Kossiakoff, A., Noinaj, N.(2023) Structure 31: 595-606.e5
- PubMed: 36977410 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2023.03.002
- Primary Citation Related Structures:
8DN6, 8DN7 - PubMed Abstract:
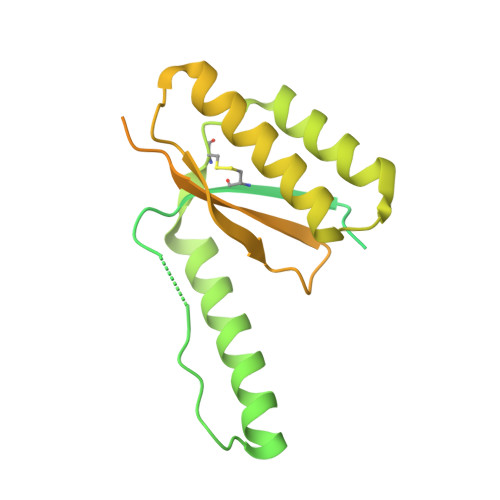
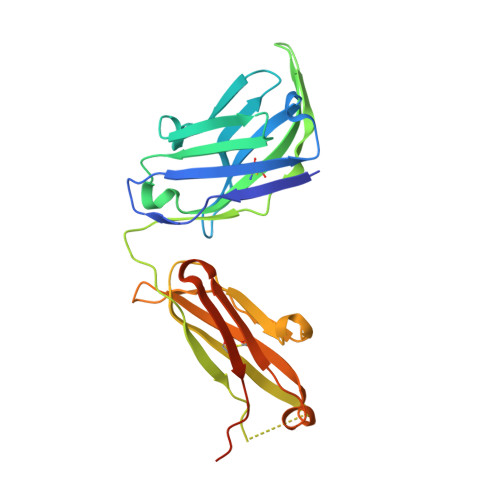
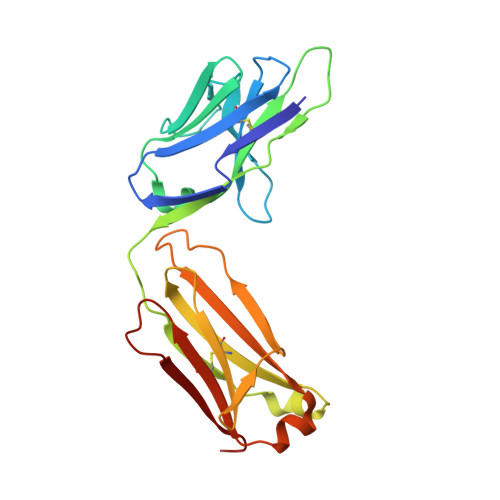
Roughly 95% of the proteins that make up the chloroplast must be imported from the cytoplasm. The machinery responsible for the translocation of these cargo proteins is called the translocon at the outer membrane of chloroplast (TOC). The TOC core consists of three proteins, Toc34, Toc75, and Toc159; no high-resolution structure has been solved of fully assembled TOC from plants. Efforts toward determining the structure of the TOC have been hindered almost entirely by difficulties in producing sufficient yields for structural studies. In this study, we introduce an innovative method that utilizes synthetic antigen binding fragments (sABs) to isolate TOC directly from wild-type plant biomass including A. thaliana and P. sativum. Binding between the sABs and the POTRA domains was characterized by size-exclusion chromatography coupled with small-angle X-ray scattering (SEC-SAXS), X-ray crystallography, and isothermal titration calorimetry. We also demonstrate the isolation of the TOC from P. sativum, laying the framework for large-scale isolation and purification of TOC for functional and structural studies.
- Department of Biological Sciences, Purdue University, West Lafayette, IN, USA.
Organizational Affiliation: