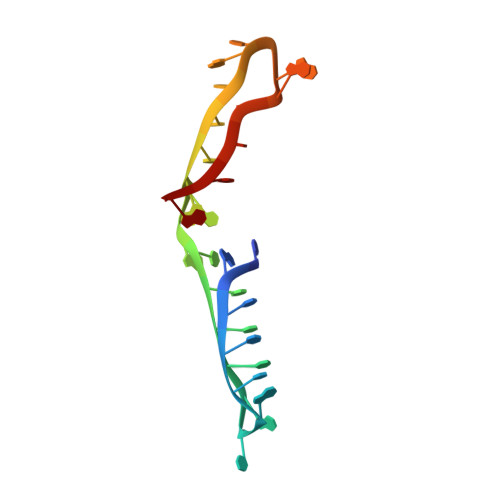
Crystal Structure of an i-Motif from the HRAS Oncogene Promoter.
Li, K.S., Jordan, D., Lin, L.Y., McCarthy, S.E., Schneekloth Jr., J.S., Yatsunyk, L.A.(2023) Angew Chem Int Ed Engl 62: e202301666-e202301666
- PubMed: 36995904 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202301666
- Primary Citation Related Structures:
8CXF, 8DHC - PubMed Abstract:
An i-motif is a non-canonical DNA structure implicated in gene regulation and linked to cancers. The C-rich strand of the HRAS oncogene, 5'-CGCCCGTGCCCTGCGCCCGCAACCCGA-3' (herein referred to as iHRAS), forms an i-motif in vitro but its exact structure was unknown. HRAS is a member of the RAS proto-oncogene family. About 19 % of US cancer patients carry mutations in RAS genes. We solved the structure of iHRAS at 1.77 Å resolution. The structure reveals that iHRAS folds into a double hairpin. The two double hairpins associate in an antiparallel fashion, forming an i-motif dimer capped by two loops on each end and linked by a connecting region. Six C-C + base pairs form each i-motif core, and the core regions are extended by a G-G base pair and a cytosine stacking. Extensive canonical and non-canonical base pairing and stacking stabilizes the connecting region and loops. The iHRAS structure is the first atomic resolution structure of an i-motif from a human oncogene. This structure sheds light on i-motifs folding and function in the cell.
- Department Chemistry and Biochemistry, Swarthmore College, 500 College Ave, Swarthmore, PA 19081, USA.
Organizational Affiliation: