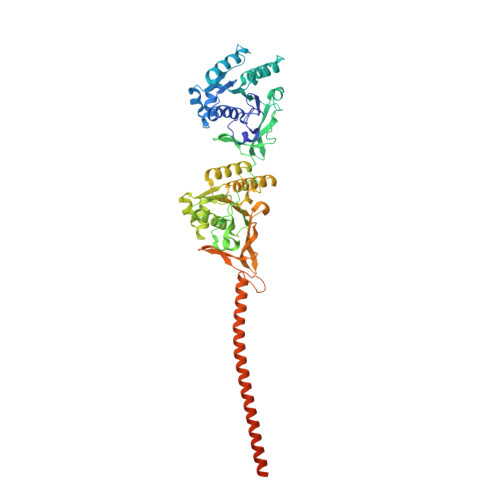
From primordial clocks to circadian oscillators.
Pitsawong, W., Padua, R.A.P., Grant, T., Hoemberger, M., Otten, R., Bradshaw, N., Grigorieff, N., Kern, D.(2023) Nature 616: 183-189
- PubMed: 36949197 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-023-05836-9
- Primary Citation Related Structures:
8DB3, 8DBA, 8FWI, 8FWJ - PubMed Abstract:
Circadian rhythms play an essential part in many biological processes, and only three prokaryotic proteins are required to constitute a true post-translational circadian oscillator 1 . The evolutionary history of the three Kai proteins indicates that KaiC is the oldest member and a central component of the clock 2 . Subsequent additions of KaiB and KaiA regulate the phosphorylation state of KaiC for time synchronization. The canonical KaiABC system in cyanobacteria is well understood 3-6 , but little is known about more ancient systems that only possess KaiBC. However, there are reports that they might exhibit a basic, hourglass-like timekeeping mechanism 7-9 . Here we investigate the primordial circadian clock in Rhodobacter sphaeroides, which contains only KaiBC, to elucidate its inner workings despite missing KaiA. Using a combination of X-ray crystallography and cryogenic electron microscopy, we find a new dodecameric fold for KaiC, in which two hexamers are held together by a coiled-coil bundle of 12 helices. This interaction is formed by the carboxy-terminal extension of KaiC and serves as an ancient regulatory moiety that is later superseded by KaiA. A coiled-coil register shift between daytime and night-time conformations is connected to phosphorylation sites through a long-range allosteric network that spans over 140 Å. Our kinetic data identify the difference in the ATP-to-ADP ratio between day and night as the environmental cue that drives the clock. They also unravel mechanistic details that shed light on the evolution of self-sustained oscillators.
- Howard Hughes Medical Institute and Department of Biochemistry, Brandeis University, Waltham, MA, USA.
Organizational Affiliation: