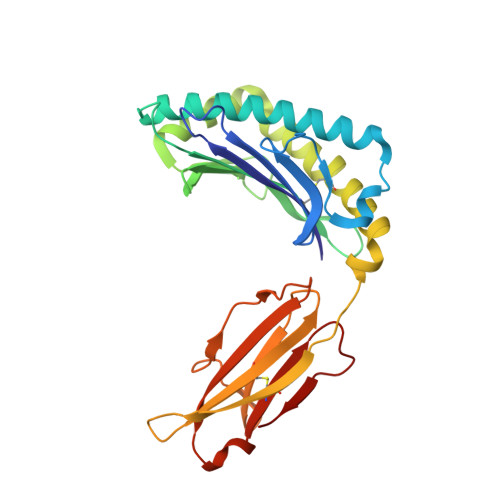
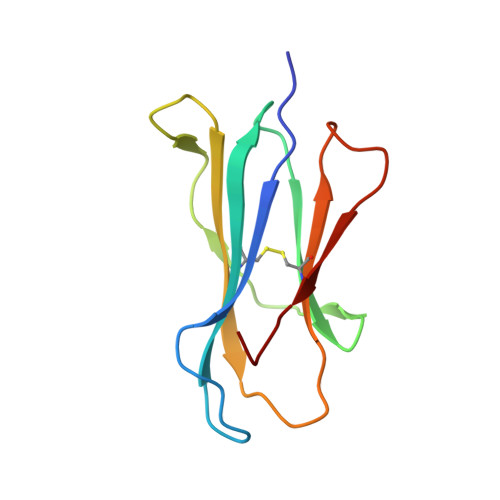
Structural and physical features that distinguish tumor-controlling from inactive cancer neoepitopes.
Custodio, J.M., Ayres, C.M., Rosales, T.J., Brambley, C.A., Arbuiso, A.G., Landau, L.M., Keller, G.L.J., Srivastava, P.K., Baker, B.M.(2023) Proc Natl Acad Sci U S A 120: e2312057120-e2312057120
- PubMed: 38085776 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2312057120
- Primary Citation Related Structures:
8D5E, 8D5F, 8D5K - PubMed Abstract:
Neoepitopes arising from amino acid substitutions due to single nucleotide polymorphisms are targets of T cell immune responses to cancer and are of significant interest in the development of cancer vaccines. However, understanding the characteristics of rare protective neoepitopes that truly control tumor growth has been a challenge, due to their scarcity as well as the challenge of verifying true, neoepitope-dependent tumor control in humans. Taking advantage of recent work in mouse models that circumvented these challenges, here, we compared the structural and physical properties of neoepitopes that range from fully protective to immunologically inactive. As neoepitopes are derived from self-peptides that can induce immune tolerance, we studied not only how the various neoepitopes differ from each other but also from their wild-type counterparts. We identified multiple features associated with protection, including features that describe how neoepitopes differ from self as well as features associated with recognition by diverse T cell receptor repertoires. We demonstrate both the promise and limitations of neoepitope structural analysis and predictive modeling and illustrate important aspects that can be incorporated into neoepitope prediction pipelines.
- Department of Chemistry and Biochemistry and the Harper Cancer Research Institute, University of Notre Dame, Notre Dame, IN 46556.
Organizational Affiliation: