Structure-Reactivity Studies of 2-Sulfonylpyrimidines Allow Selective Protein Arylation.
Pichon, M.M., Drelinkiewicz, D., Lozano, D., Moraru, R., Hayward, L.J., Jones, M., McCoy, M.A., Allstrum-Graves, S., Balourdas, D.I., Joerger, A.C., Whitby, R.J., Goldup, S.M., Wells, N., Langley, G.J., Herniman, J.M., Baud, M.G.J.(2023) Bioconjug Chem 34: 1679-1687
- PubMed: 37657082 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.bioconjchem.3c00322
- Primary Citation Related Structures:
8CG7 - PubMed Abstract:
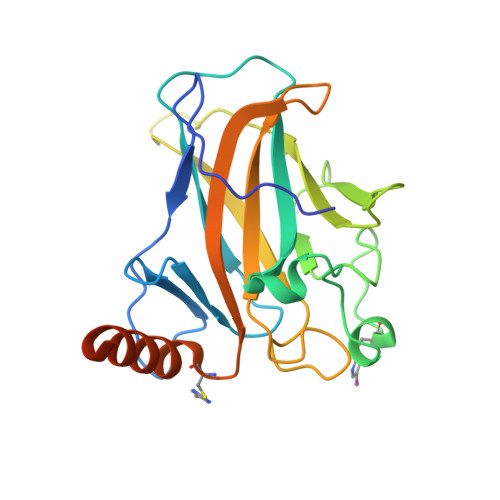
Protein arylation has attracted much attention for developing new classes of bioconjugates with improved properties. Here, we have evaluated 2-sulfonylpyrimidines as covalent warheads for the mild, chemoselective, and metal free cysteine S -arylation. 2-Sulfonylpyrimidines react rapidly with cysteine, resulting in stable S -heteroarylated adducts at neutral pH. Fine tuning the heterocyclic core and exocyclic leaving group allowed predictable S N Ar reactivity in vitro , covering >9 orders of magnitude. Finally, we achieved fast chemo- and regiospecific arylation of a mutant p53 protein and confirmed arylation sites by protein X-ray crystallography. Hence, we report the first example of a protein site specifically S -arylated with iodo-aromatic motifs. Overall, this study provides the most comprehensive structure-reactivity relationship to date on heteroaryl sulfones and highlights 2-sulfonylpyrimidine as a synthetically tractable and protein compatible covalent motif for targeting reactive cysteines, expanding the arsenal of tunable warheads for modern covalent ligand discovery.
- School of Chemistry, University of Southampton, Highfield, SO17 1BJ Southampton, United Kingdom.
Organizational Affiliation: