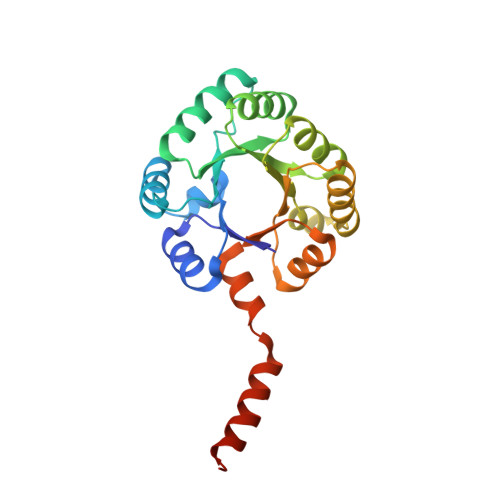
Structure and mechanism of sulfofructose transaldolase, a key enzyme in sulfoquinovose metabolism.
Snow, A.J.D., Sharma, M., Abayakoon, P., Williams, S.J., Blaza, J.N., Davies, G.J.(2023) Structure 31: 244
- PubMed: 36805128 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2023.01.010
- Primary Citation Related Structures:
8BC2, 8BC3, 8BC4, 8C4I - PubMed Abstract:
Sulfoquinovose (SQ) is a key component of plant sulfolipids (sulfoquinovosyl diacylglycerols) and a major environmental reservoir of biological sulfur. Breakdown of SQ is achieved by bacteria through the pathways of sulfoglycolysis. The sulfoglycolytic sulfofructose transaldolase (sulfo-SFT) pathway is used by gut-resident firmicutes and soil saprophytes. After isomerization of SQ to sulfofructose (SF), the namesake enzyme catalyzes the transaldol reaction of SF transferring dihydroxyacetone to 3C/4C acceptors to give sulfolactaldehyde and fructose-6-phosphate or sedoheptulose-7-phosphate. We report the 3D cryo-EM structure of SF transaldolase from Bacillus megaterium in apo and ligand bound forms, revealing a decameric structure formed from two pentameric rings of the protomer. We demonstrate a covalent "Schiff base" intermediate formed by reaction of SF with Lys89 within a conserved Asp-Lys-Glu catalytic triad and defined by an Arg-Trp-Arg sulfonate recognition triad. The structural characterization of the signature enzyme of the sulfo-SFT pathway provides key insights into molecular recognition of the sulfonate group of sulfosugars.
- Department of Chemistry, York Structural Biology Laboratory, York YO10 5DD, UK.
Organizational Affiliation: