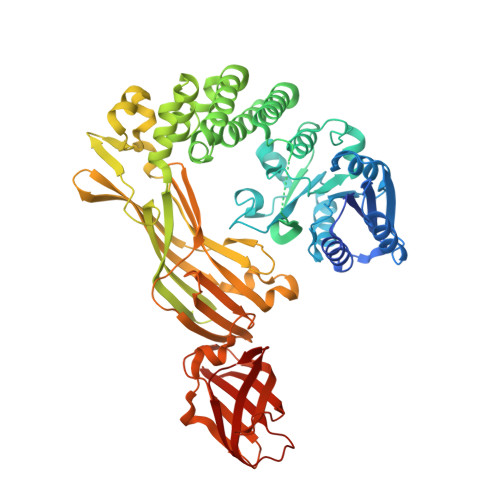
Molecular properties of the RmlT wall teichoic acid rhamnosyltransferase that modulates virulence in Listeria monocytogenes.
Monteiro, R., Cereija, T.B., Pombinho, R., Voskuilen, T., Codee, J.D.C., Sousa, S., Morais-Cabral, J.H., Cabanes, D.(2025) Nat Commun 16: 24-24
- PubMed: 39746981 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-55360-1
- Primary Citation Related Structures:
8BZ4, 8BZ5, 8BZ6, 8BZ7, 8BZ8, 9GZJ, 9GZK - PubMed Abstract:
Wall teichoic acids (WTAs) from the major Gram-positive foodborne pathogen Listeria monocytogenes are peptidoglycan-associated glycopolymers decorated by monosaccharides that, while not essential for bacterial growth, are required for bacterial virulence and resistance to antimicrobials. Here we report the structure and function of a bacterial WTAs rhamnosyltransferase, RmlT, strictly required for L. monocytogenes WTAs rhamnosylation. In particular, we demonstrated that RmlT transfers rhamnose from dTDP-L-rhamnose to naked WTAs, and that specificity towards TDP-rhamnose is not determined by its binding affinity. Structures of RmlT with and without its substrates showed that this enzyme is a dimer, revealed the residues responsible for interaction with the substrates and that the catalytic residue pre-orients the acceptor substrate towards the nucleophilic attack to the sugar. Additionally, the structures provided indications for two potential interaction pathways for the long WTAs on the surface of RmlT. Finally, we confirmed that WTAs glycosyltransferases are promising targets for next-generation strategies against Gram-positive pathogens by showing that inactivation of the RmlT catalytic activity results in a decreased infection in vivo.
- i3S - Instituto de Investigação e Inovação em Saúde, Universidade do Porto, Porto, Portugal. ricardo.monteiro@i3s.up.pt.
Organizational Affiliation: