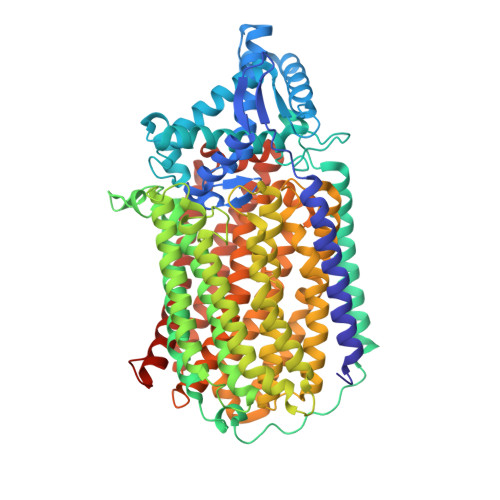
A 2.2 angstrom cryoEM structure of a quinol-dependent NO Reductase shows close similarity to respiratory oxidases.
Flynn, A.J., Antonyuk, S.V., Eady, R.R., Muench, S.P., Hasnain, S.S.(2023) Nat Commun 14: 3416-3416
- PubMed: 37296134 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-39140-x
- Primary Citation Related Structures:
8BGW - PubMed Abstract:
Quinol-dependent nitric oxide reductases (qNORs) are considered members of the respiratory heme-copper oxidase superfamily, are unique to bacteria, and are commonly found in pathogenic bacteria where they play a role in combating the host immune response. qNORs are also essential enzymes in the denitrification pathway, catalysing the reduction of nitric oxide to nitrous oxide. Here, we determine a 2.2 Å cryoEM structure of qNOR from Alcaligenes xylosoxidans, an opportunistic pathogen and a denitrifying bacterium of importance in the nitrogen cycle. This high-resolution structure provides insight into electron, substrate, and proton pathways, and provides evidence that the quinol binding site not only contains the conserved His and Asp residues but also possesses a critical Arg (Arg720) observed in cytochrome bo 3 , a respiratory quinol oxidase.
- School of Biomedical Sciences, Faculty of Biological Sciences, University of Leeds, Leeds, LS2 9JT, UK.
Organizational Affiliation: