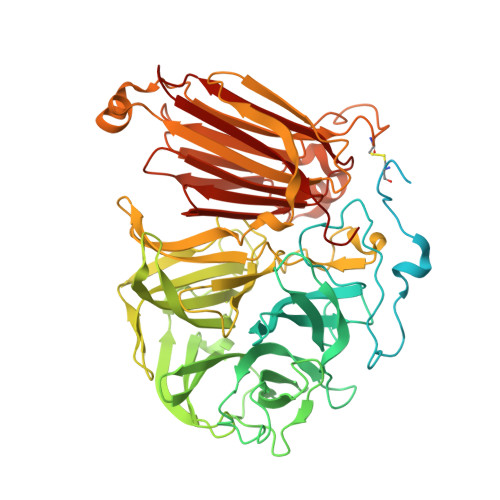
Insights into the Structure of the Highly Glycosylated Ffase from Rhodotorula dairenensis Enhance Its Biotechnological Potential.
Jimenez-Ortega, E., Narmontaite, E., Gonzalez-Perez, B., Plou, F.J., Fernandez-Lobato, M., Sanz-Aparicio, J.(2022) Int J Mol Sci 23
- PubMed: 36499311 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms232314981
- Primary Citation Related Structures:
8BEQ, 8BES, 8BET, 8BEU - PubMed Abstract:
Rhodotorula dairenensis β-fructofuranosidase is a highly glycosylated enzyme with broad substrate specificity that catalyzes the synthesis of 6-kestose and a mixture of the three series of fructooligosaccharides (FOS), fructosylating a variety of carbohydrates and other molecules as alditols. We report here its three-dimensional structure, showing the expected bimodular arrangement and also a unique long elongation at its N-terminus containing extensive O-glycosylation sites that form a peculiar arrangement with a protruding loop within the dimer. This region is not required for activity but could provide a molecular tool to target the dimeric protein to its receptor cellular compartment in the yeast. A truncated inactivated form was used to obtain complexes with fructose, sucrose and raffinose, and a Bis-Tris molecule was trapped, mimicking a putative acceptor substrate. The crystal structure of the complexes reveals the major traits of the active site, with Asn387 controlling the substrate binding mode. Relevant residues were selected for mutagenesis, the variants being biochemically characterized through their hydrolytic and transfructosylating activity. All changes decrease the hydrolytic efficiency against sucrose, proving their key role in the activity. Moreover, some of the generated variants exhibit redesigned transfructosylating specificity, which may be used for biotechnological purposes to produce novel fructosyl-derivatives.
- Department of Crystallography and Structural Biology, Institute of Physical-Chemistry Rocasolano, CSIC, 28006 Madrid, Spain.
Organizational Affiliation: