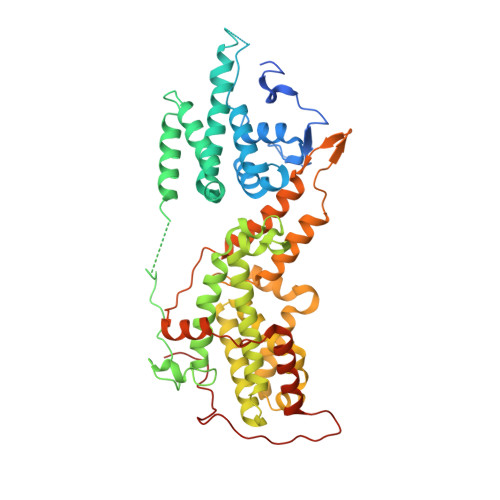
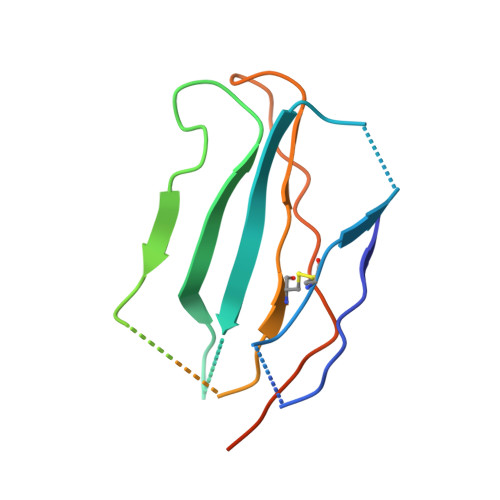
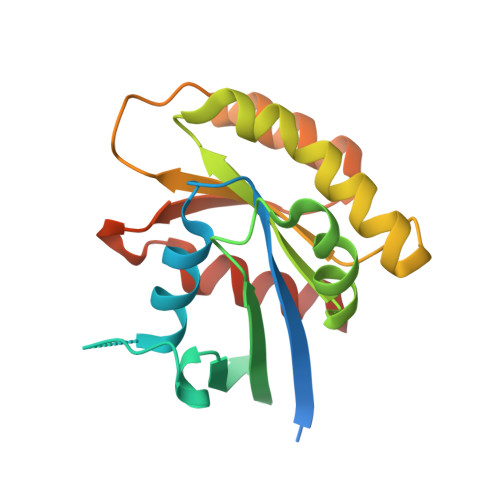
Allosteric nanobodies to study the interactions between SOS1 and RAS.
Fischer, B., Uchanski, T., Sheryazdanova, A., Gonzalez, S., Volkov, A.N., Brosens, E., Zogg, T., Kalichuk, V., Ballet, S., Versees, W., Sablina, A.A., Pardon, E., Wohlkonig, A., Steyaert, J.(2024) Nat Commun 15: 6214-6214
- PubMed: 39043660 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-50349-2
- Primary Citation Related Structures:
8BE2, 8BE3, 8BE4, 8BE5 - PubMed Abstract:
Protein-protein interactions (PPIs) are central in cell metabolism but research tools for the structural and functional characterization of these PPIs are often missing. Here we introduce broadly applicable immunization (Cross-link PPIs and immunize llamas, ChILL) and selection strategies (Display and co-selection, DisCO) for the discovery of diverse nanobodies that either stabilize or disrupt PPIs in a single experiment. We apply ChILL and DisCO to identify competitive, connective, or fully allosteric nanobodies that inhibit or facilitate the formation of the SOS1•RAS complex and modulate the nucleotide exchange rate on this pivotal GTPase in vitro as well as RAS signalling in cellulo. One of these connective nanobodies fills a cavity that was previously identified as the binding pocket for a series of therapeutic lead compounds. The long complementarity-determining region (CDR3) that penetrates this binding pocket serves as pharmacophore for extending the repertoire of potential leads.
- Université de Bordeaux, CNRS, Bordeaux INP, CBMN, UMR 5248, Pessac, France.
Organizational Affiliation: