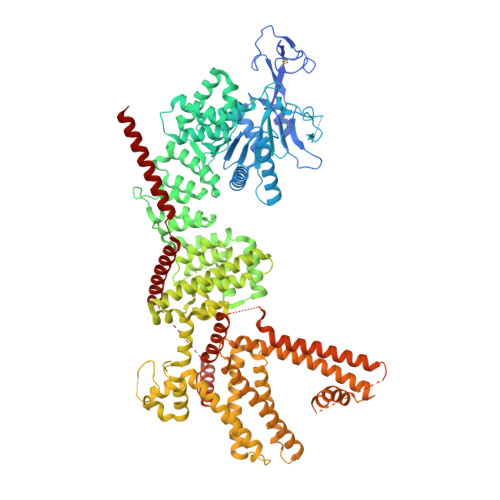
Structure of human TRPM8 channel.
Palchevskyi, S., Czarnocki-Cieciura, M., Vistoli, G., Gervasoni, S., Nowak, E., Beccari, A.R., Nowotny, M., Talarico, C.(2023) Commun Biol 6: 1065-1065
- PubMed: 37857704 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-023-05425-6
- Primary Citation Related Structures:
8BDC - PubMed Abstract:
TRPM8 is a non-selective cation channel permeable to both monovalent and divalent cations that is activated by multiple factors, such as temperature, voltage, pressure, and changes in osmolality. It is a therapeutic target for anticancer drug development, and its modulators can be utilized for several pathological conditions. Here, we present a cryo-electron microscopy structure of a human TRPM8 channel in the closed state that was solved at 2.7 Å resolution. Our structure comprises the most complete model of the N-terminal pre-melastatin homology region. We also visualized several lipids that are bound by the protein and modeled how the human channel interacts with icilin. Analyses of pore helices in available TRPM structures showed that all these structures can be grouped into different closed, desensitized and open state conformations based on the register of the pore helix S6 which positions particular amino acid residues at the channel constriction.
- Laboratory of Protein Structure, International Institute of Molecular and Cell Biology in Warsaw, 02-109, Warsaw, Poland.
Organizational Affiliation: