Pathogenic antibody response to glucose-6-phosphate isomerase targets a modified epitope uniquely exposed on joint cartilage.
Li, T., Ge, C., Kramer, A., Sareila, O., Leu Agelii, M., Johansson, L., Forslind, K., Lonnblom, E., Yang, M., Xu, B., Li, Q., Cheng, L., Bergstrom, G., Fernandez, G., Kastbom, A., Rantapaa-Dahlqvist, S., Gjertsson, I., Holmdahl, R.(2023) Ann Rheum Dis 82: 799-808
- PubMed: 36858822 Search on PubMed
- DOI: https://doi.org/10.1136/ard-2022-223633
- Primary Citation Related Structures:
8BBH - PubMed Abstract:
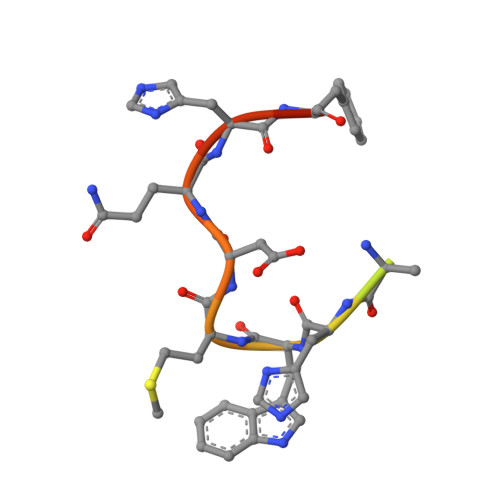
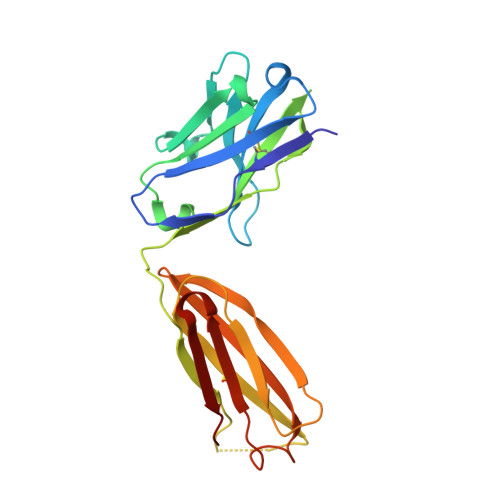
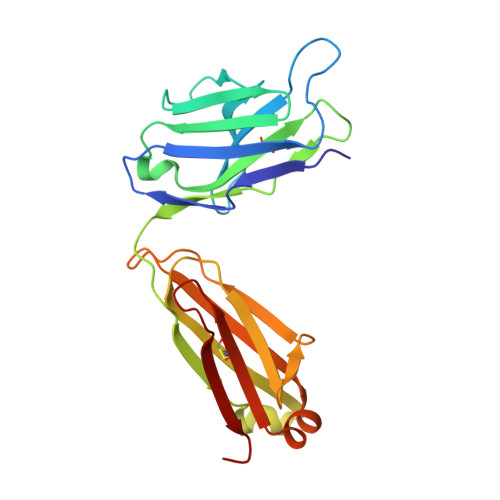
To identify the arthritogenic B cell epitopes of glucose-6-phosphate isomerase (GPI) and their association with rheumatoid arthritis (RA). IgG response towards a library of GPI peptides in patients with early RA, pre-symptomatic individuals and population controls, as well as in mice, were tested by bead-based multiplex immunoassays and ELISA. Monoclonal IgG were generated, and the binding specificity and affinity were determined by ELISA, gel size exclusion chromatography, surface plasma resonance and X-ray crystallography. Arthritogenicity was investigated by passive transfer experiments. Antigen-specific B cells were identified by peptide tetramer staining. Peptide GPI 293-307 was the dominant B cell epitope in K/BxN and GPI-immunised mice. We could detect B cells and low levels of IgM antibodies binding the GPI 293-307 epitopes, and high affinity anti-GPI 293-307 IgG antibodies already 7 days after GPI immunisation, immediately before arthritis onset. Transfer of anti-GPI 293-307 IgG antibodies induced arthritis in mice. Moreover, anti-GPI 293-307 IgG antibodies were more frequent in individuals prior to RA onset (19%) than in controls (7.5%). GPI 293-307 -specific antibodies were associated with radiographic joint damage. Crystal structures of the Fab-peptide complex revealed that this epitope is not exposed in native GPI but requires conformational change of the protein in inflamed joint for effective recognition by anti-GPI 293-307 antibodies. We have identified the major pathogenic B cell epitope of the RA-associated autoantigen GPI, at position 293-307, exposed only on structurally modified GPI on the cartilage surface. B cells to this neo-epitope escape tolerance and could potentially play a role in the pathogenesis of RA.
- Section of Medical Inflammation Research, Department of Medical Biochemistry and Biophysics, Karolinska Institute, Stockholm, Sweden.
Organizational Affiliation: