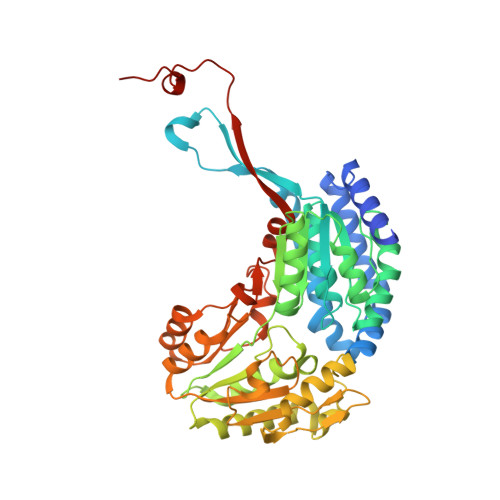
Characterization of human oxidoreductases involved in aldehyde odorant metabolism.
Boichot, V., Menetrier, F., Saliou, J.M., Lirussi, F., Canon, F., Folia, M., Heydel, J.M., Hummel, T., Menzel, S., Steinke, M., Hackenberg, S., Schwartz, M., Neiers, F.(2023) Sci Rep 13: 4876-4876
- PubMed: 36966166 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-023-31769-4
- Primary Citation Related Structures:
8BB8 - PubMed Abstract:
Oxidoreductases are major enzymes of xenobiotic metabolism. Consequently, they are essential in the chemoprotection of the human body. Many xenobiotic metabolism enzymes have been shown to be involved in chemosensory tissue protection. Among them, some were additionally shown to be involved in chemosensory perception, acting in signal termination as well as in the generation of metabolites that change the activation pattern of chemosensory receptors. Oxidoreductases, especially aldehyde dehydrogenases and aldo-keto reductases, are the first barrier against aldehyde compounds, which include numerous odorants. Using a mass spectrometry approach, we characterized the most highly expressed members of these families in the human nasal mucus sampled in the olfactory vicinity. Their expression was also demonstrated using immunohistochemistry in human epitheliums sampled in the olfactory vicinity. Recombinant enzymes corresponding to three highly expressed human oxidoreductases (ALDH1A1, ALDH3A1, AKR1B10) were used to demonstrate the high enzymatic activity of these enzymes toward aldehyde odorants. The structure‒function relationship set based on the enzymatic parameters characterization of a series of aldehyde odorant compounds was supported by the X-ray structure resolution of human ALDH3A1 in complex with octanal.
- Flavour Perception: Molecular Mechanisms (Flavours), INRAE, CNRS, Institut Agro, Université de Bourgogne Franche-Comté, Dijon, France.
Organizational Affiliation: