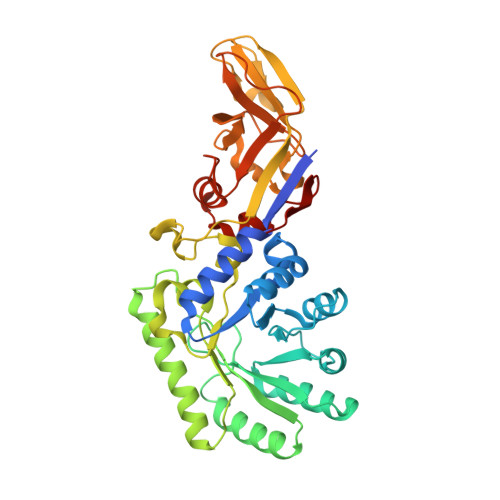
Structure of the d-Cycloserine-Resistant Variant D322N of Alanine Racemase from Mycobacterium tuberculosis .
de Chiara, C., Prosser, G.A., Ogrodowicz, R., de Carvalho, L.P.S.(2023) ACS Bio Med Chem Au 3: 233-239
- PubMed: 37363078 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsbiomedchemau.2c00074
- Primary Citation Related Structures:
8AHW, 8B8H - PubMed Abstract:
Alanine racemase (Alr) is a pyridoxal 5'-phosphate-dependent enzyme that catalyzes the racemization of l-alanine to d-alanine. Alr is one of the two targets of the broad-spectrum antibiotic d-cycloserine (DCS), a structural analogue of d-alanine. Despite being an essential component of regimens used to treat multi- and extensively drug-resistant tuberculosis for almost seven decades, resistance to DCS has not been observed in patients. We previously demonstrated that DCS evades resistance due to an ultralow rate of emergence of mutations. Yet, we identified a single polymorphism (converting Asp322 to Asn) in the alr gene, which arose in 8 out of 11 independent variants identified and that confers resistance. Here, we present the crystal structure of the Alr variant D322N in both the free and DCS-inactivated forms and the characterization of its DCS inactivation mechanism by UV-visible and fluorescence spectroscopy. Comparison of these results with those obtained with wild-type Alr reveals the structural basis of the 240-fold reduced inhibition observed in Alr D322N.
- Mycobacterial Metabolism and Antibiotic Research Laboratory, The Francis Crick Institute, London, NW1 1AT, United Kingdom.
Organizational Affiliation: