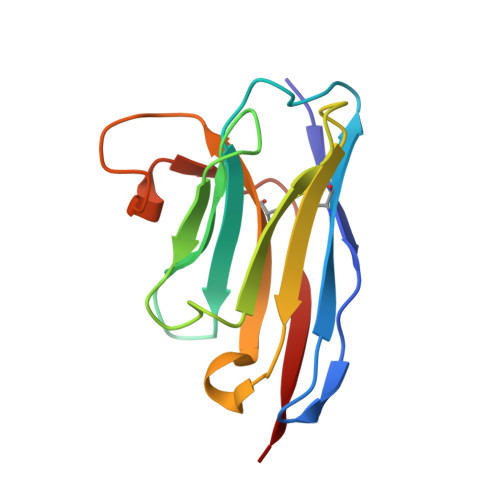
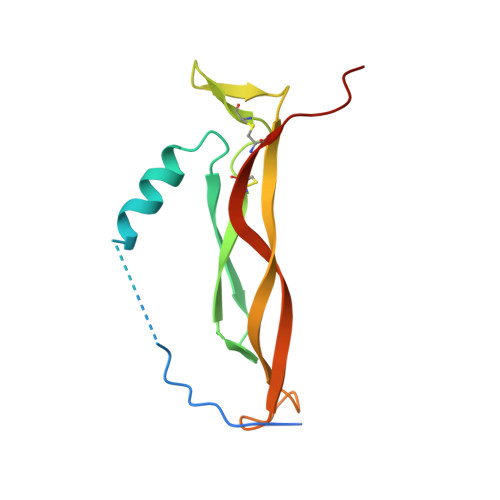
Two Epitope Regions Revealed in the Complex of IL-17A and Anti-IL-17A V H H Domain.
Kostareva, O., Svoeglazova, A., Kolyadenko, I., Nikulin, A., Evdokimov, S., Dzhus, U., Gabdulkhakov, A., Tishchenko, S.(2022) Int J Mol Sci 23
- PubMed: 36499233 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms232314904
- Primary Citation Related Structures:
8B7W - PubMed Abstract:
Interleukin-17 (IL-17) is a cytokine produced by the Th17 cells. It is involved in chronic inflammation in patients with autoimmune diseases, such as rheumatoid arthritis, systemic lupus erythematosus, multiple sclerosis, and psoriasis. The antibodies targeting IL-17 and/or IL-17R are therapy tools for these diseases. Netakimab is an IL-17A-specific antibody containing a Lama glama V H H derivative domain and a VL variable domain. We have determined the crystal structure of the IL-17A-specific V H H domain in complex with IL-17A at 2.85 Å resolution. Certain amino acid residues of the three complementary-determining regions of the V H H domain form a network of solvent-inaccessible hydrogen bonds with two epitope regions of IL-17A. The β-turn of IL-17A, which forms the so-called epitope-1, appears to be the main region of IL-17A interaction with the antibody. Contacts formed by the IL-17A mobile C-terminal region residues (epitope-2) further stabilize the antibody-antigen complex.
- Institute of Protein Research, Russian Academy of Sciences, Institutskaya, 4, 142290 Pushchino, Russia.
Organizational Affiliation: