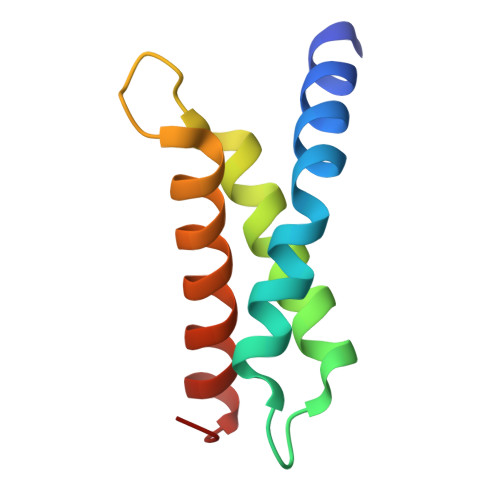
Structural Analysis of Bacillus subtilis Sigma Factors.
Collins, K.M., Evans, N.J., Torpey, J.H., Harris, J.M., Haynes, B.A., Camp, A.H., Isaacson, R.L.(2023) Microorganisms 11
- PubMed: 37110501 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/microorganisms11041077
- Primary Citation Related Structures:
8B3Z - PubMed Abstract:
Bacteria use an array of sigma factors to regulate gene expression during different stages of their life cycles. Full-length, atomic-level structures of sigma factors have been challenging to obtain experimentally as a result of their many regions of intrinsic disorder. AlphaFold has now supplied plausible full-length models for most sigma factors. Here we discuss the current understanding of the structures and functions of sigma factors in the model organism, Bacillus subtilis , and present an X-ray crystal structure of a region of B. subtilis SigE, a sigma factor that plays a critical role in the developmental process of spore formation.
- Department of Chemistry, King's College London, Britannia House, 7 Trinity Street, London SE1 1DB, UK.
Organizational Affiliation: