Structure of YjbA in complex with ClpC N-terminal Domain
Evans, N.J., Isaacson, R.L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| UPF0736 protein B4122_0676,UPF0736 protein YjbA | 351 | Bacillus subtilis | Mutation(s): 0 Gene Names: B4122_0676, B4417_4067, Bateq7PJ16_1268, DFO69_2911, J5227_16610, SC09_Contig19orf00439, yjbA, BSU11410 | 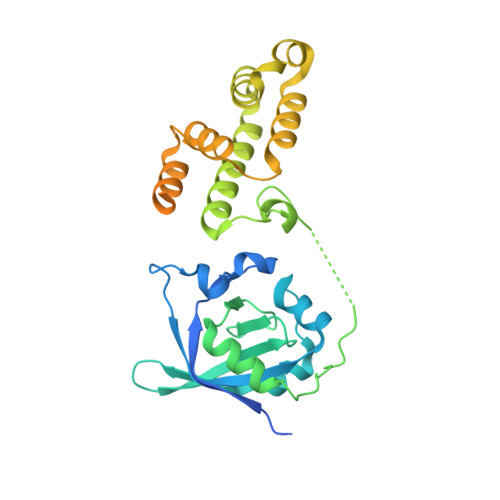 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O31597 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ATP-dependent Clp protease ATP-binding subunit ClpC / Negative regulator of tic competence clcC/mecB | B [auth Z] | 156 | Bacillus subtilis | Mutation(s): 0 Gene Names: B4417_1475 | 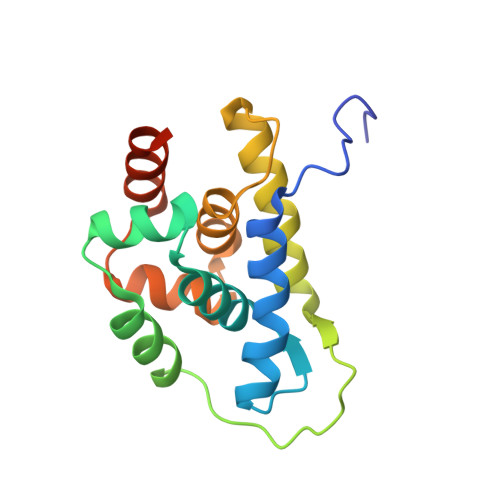 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P37571 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GOL Download:Ideal Coordinates CCD File | C [auth A] D [auth A] E [auth A] F [auth A] G [auth Z] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | I [auth Z] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 69.713 | α = 90 |
| b = 69.713 | β = 90 |
| c = 89.601 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| DIALS | data reduction |
| DIALS | data scaling |
| PHASER | phasing |
| SHELXDE | phasing |
| ARP/wARP | model building |
| BUCCANEER | model building |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Biotechnology and Biological Sciences Research Council (BBSRC) | United Kingdom | -- |