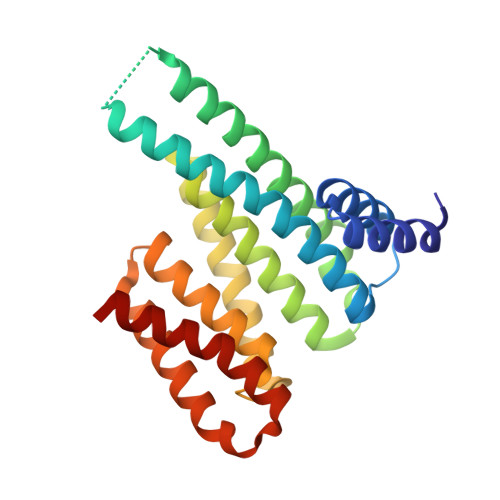
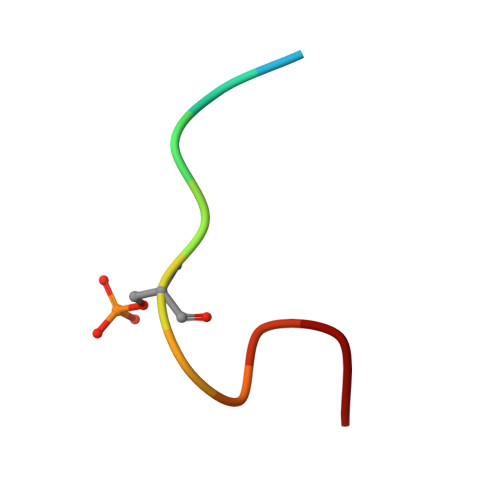
Potential 14-3-3 binding sites in sirtuins reveal extended phosphosite-recognition modes.
Weyand, M., Quast, L., Steegborn, C.(2026) Acta Crystallogr F Struct Biol Commun 82: 32-40
- PubMed: 41410196 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X25010908
- Primary Citation Related Structures:
8ANB, 8ANC - PubMed Abstract:
The adapter proteins of the 14-3-3 family modulate the activity and/or localization of their binding partners, which they capture if a generic target motif is in its phosphorylated state. Here, we report the identification of potential 14-3-3 binding sites in human sirtuin deacylases by bioinformatic analysis. We then characterize the interactions of peptides representing phosphorylation sites in sirtuin 3 (pS103) and sirtuin 1 (pS670) with 14-3-3 proteins. We further describe the crystal structures of complexes of 14-3-3σ with either of the two phosphopeptides. As a conclusion, we propose a more extended 14-3-3 binding mode on the N-terminal side of the phosphorylation site and the possibility of nongeneric motifs and conformations on the C-terminal side, still resulting in the known high binding affinity of the two partners.
- Department of Biochemistry, University of Bayreuth, Universitätsstrasse 30, 95440 Bayreuth, Germany.
Organizational Affiliation: