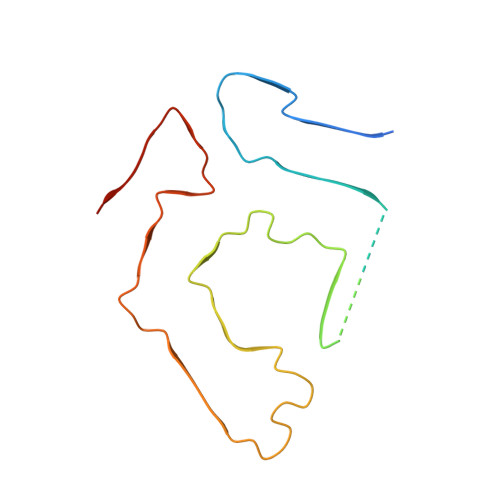
Cryo-EM structure of an ATTRwt amyloid fibril from systemic non-hereditary transthyretin amyloidosis.
Steinebrei, M., Gottwald, J., Baur, J., Rocken, C., Hegenbart, U., Schonland, S., Schmidt, M.(2022) Nat Commun 13: 6398-6398
- PubMed: 36302762 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-33591-4
- Primary Citation Related Structures:
8ADE - PubMed Abstract:
Wild type transthyretin-derived amyloid (ATTRwt) is the major component of non-hereditary transthyretin amyloidosis. Its accumulation in the heart of elderly patients is life threatening. A variety of genetic variants of transthyretin can lead to hereditary transthyretin amyloidosis, which shows different clinical symptoms, like age of onset and pattern of organ involvement. However, in the case of non-hereditary transthyretin amyloidosis ATTRwt fibril deposits are located primarily in heart tissue. In this structural study we analyzed ATTRwt amyloid fibrils from the heart of a patient with non-hereditary transthyretin amyloidosis. We present a 2.78 Å reconstructed density map of these ATTRwt fibrils using cryo electron microscopy and compare it with previously published V30M variants of ATTR fibrils extracted from heart and eye of different patients. All structures show a remarkably similar spearhead like shape in their cross section, formed by the same N- and C-terminal fragments of transthyretin with some minor differences. This demonstrates common features for ATTR fibrils despite differences in mutations and patients.
- Institute of Protein Biochemistry, Ulm University, Helmholtzstrasse 8/1, Ulm, D-89081, Germany.
Organizational Affiliation: