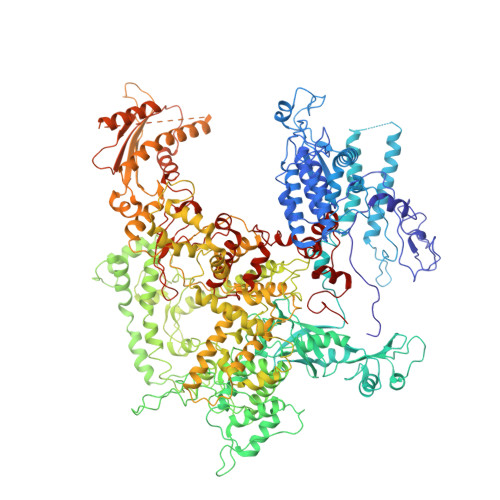
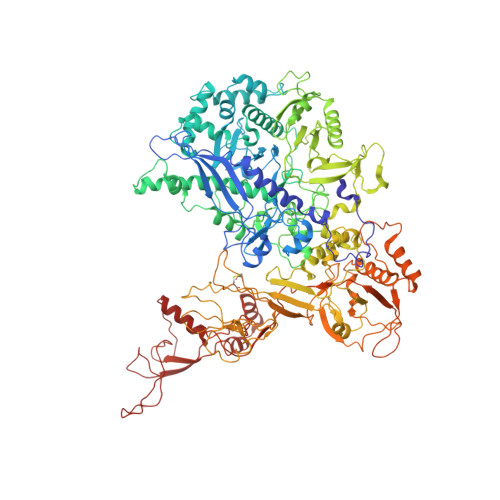
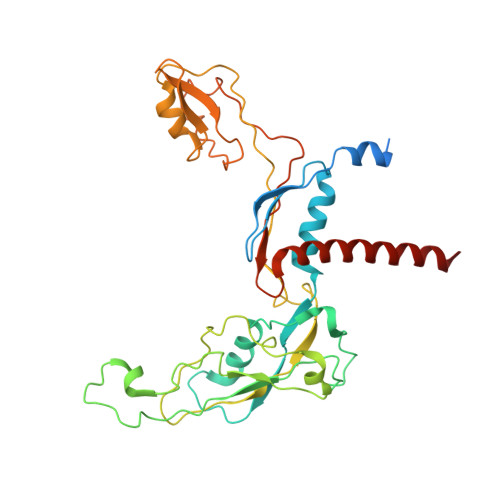
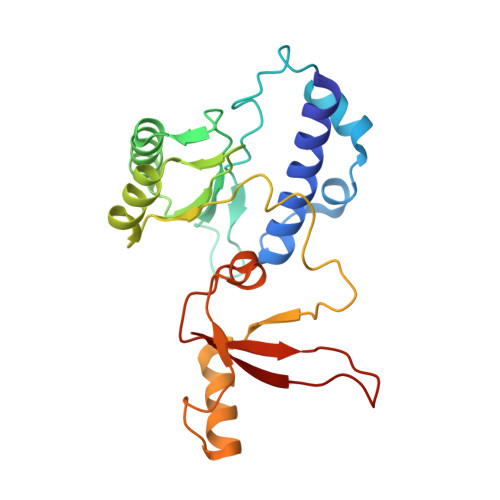
The human RNA polymerase I structure reveals an HMG-like docking domain specific to metazoans.
Daiss, J.L., Pilsl, M., Straub, K., Bleckmann, A., Hocherl, M., Heiss, F.B., Abascal-Palacios, G., Ramsay, E.P., Tluckova, K., Mars, J.C., Furtges, T., Bruckmann, A., Rudack, T., Bernecky, C., Lamour, V., Panov, K., Vannini, A., Moss, T., Engel, C.(2022) Life Sci Alliance 5
- PubMed: 36271492 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.202201568
- Primary Citation Related Structures:
8A43 - PubMed Abstract:
Transcription of the ribosomal RNA precursor by RNA polymerase (Pol) I is a major determinant of cellular growth, and dysregulation is observed in many cancer types. Here, we present the purification of human Pol I from cells carrying a genomic GFP fusion on the largest subunit allowing the structural and functional analysis of the enzyme across species. In contrast to yeast, human Pol I carries a single-subunit stalk, and in vitro transcription indicates a reduced proofreading activity. Determination of the human Pol I cryo-EM reconstruction in a close-to-native state rationalizes the effects of disease-associated mutations and uncovers an additional domain that is built into the sequence of Pol I subunit RPA1. This "dock II" domain resembles a truncated HMG box incapable of DNA binding which may serve as a downstream transcription factor-binding platform in metazoans. Biochemical analysis, in situ modelling, and ChIP data indicate that Topoisomerase 2a can be recruited to Pol I via the domain and cooperates with the HMG box domain-containing factor UBF. These adaptations of the metazoan Pol I transcription system may allow efficient release of positive DNA supercoils accumulating downstream of the transcription bubble.
- Regensburg Center for Biochemistry, University of Regensburg, Regensburg, Germany.
Organizational Affiliation: