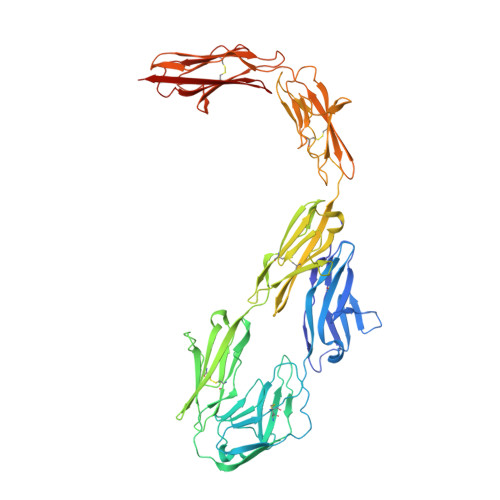
Contactin 2 homophilic adhesion structure and conformational plasticity.
Chataigner, L.M.P., Tharichen, L., Beugelink, J.W., Granneman, J.C.M., Mokiem, N.J., Snijder, J., Forster, F., Janssen, B.J.C.(2024) Structure 32: 60
- PubMed: 37992710 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2023.10.012
- Primary Citation Related Structures:
8A0Y - PubMed Abstract:
The cell-surface attached glycoprotein contactin 2 is ubiquitously expressed in the nervous system and mediates homotypic cell-cell interactions to organize cell guidance, differentiation, and adhesion. Contactin 2 consists of six Ig and four fibronectin type III domains (FnIII) of which the first four Ig domains form a horseshoe structure important for homodimerization and oligomerization. Here we report the crystal structure of the six-domain contactin 2 Ig1-6 and show that the Ig5-Ig6 combination is oriented away from the horseshoe with flexion in interdomain connections. Two distinct dimer states, through Ig1-Ig2 and Ig3-Ig6 interactions, together allow formation of larger oligomers. Combined size exclusion chromatography with multiangle light scattering (SEC-MALS), small-angle X-ray scattering (SAXS) and native MS analysis indicates contactin 2 Ig1-6 oligomerizes in a glycan dependent manner. SAXS and negative-stain electron microscopy reveals inherent plasticity of the contactin 2 full-ectodomain. The combination of intermolecular binding sites and ectodomain plasticity explains how contactin 2 can function as a homotypic adhesion molecule in diverse intercellular environments.
- Structural Biochemistry, Bijvoet Centre for Biomolecular Research, Faculty of Science, Utrecht University, Universiteitsweg 99, Utrecht 3584 CG, the Netherlands.
Organizational Affiliation: