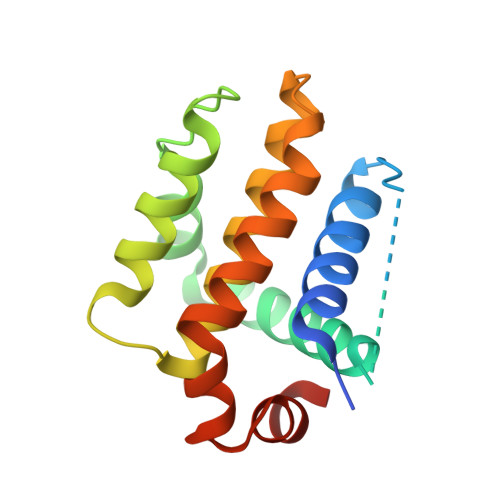
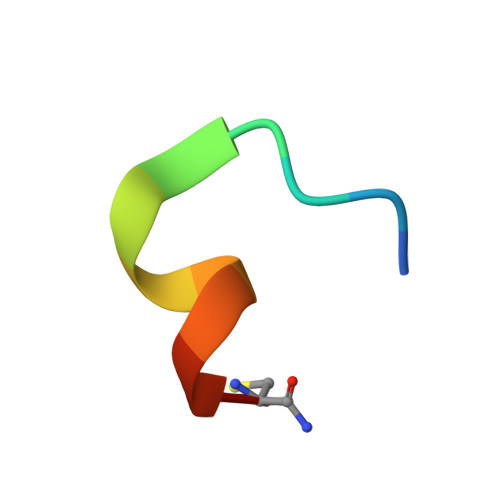
Cyclic peptides discriminate BCL-2 and its clinical mutants from BCL-X L by engaging a single-residue discrepancy.
Li, F., Liu, J., Liu, C., Liu, Z., Peng, X., Huang, Y., Chen, X., Sun, X., Wang, S., Chen, W., Xiong, D., Diao, X., Wang, S., Zhuang, J., Wu, C., Wu, D.(2024) Nat Commun 15: 1476-1476
- PubMed: 38368459 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-45848-1
- Primary Citation Related Structures:
7Y8D, 7Y90, 7Y99, 7YA5, 7YAA, 7YB7 - PubMed Abstract:
Overexpressed pro-survival B-cell lymphoma-2 (BCL-2) family proteins BCL-2 and BCL-X L can render tumor cells malignant. Leukemia drug venetoclax is currently the only approved selective BCL-2 inhibitor. However, its application has led to an emergence of resistant mutations, calling for drugs with an innovative mechanism of action. Herein we present cyclic peptides (CPs) with nanomolar-level binding affinities to BCL-2 or BCL-X L , and further reveal the structural and functional mechanisms of how these CPs target two proteins in a fashion that is remarkably different from traditional small-molecule inhibitors. In addition, these CPs can bind to the venetoclax-resistant clinical BCL-2 mutants with similar affinities as to the wild-type protein. Furthermore, we identify a single-residue discrepancy between BCL-2 D111 and BCL-X L A104 as a molecular "switch" that can differently engage CPs. Our study suggests that CPs may inhibit BCL-2 or BCL-X L by delicately modulating protein-protein interactions, potentially benefiting the development of next-generation therapeutics.
- Helmholtz International Lab, State Key Laboratory of Microbial Technology, Shandong University, Qingdao, 266237, China. lifengwei@sdu.edu.cn.
Organizational Affiliation: