Novel bispecific human antibody platform specifically targeting a fully open spike conformation potently neutralizes multiple SARS-CoV-2 variants.
Kim, J.W., Heo, K., Kim, H.J., Yoo, Y., Cho, H.S., Jang, H.J., Lee, H.Y., Ko, I.Y., Woo, J.R., Cho, Y.B., Lee, J.H., Yang, H.R., Shin, H.G., Choi, H.L., Hwang, K., Kim, S., Kim, H., Chun, K., Lee, S.(2023) Antiviral Res 212: 105576-105576
- PubMed: 36870394 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.antiviral.2023.105576
- Primary Citation Related Structures:
7Y6K - PubMed Abstract:
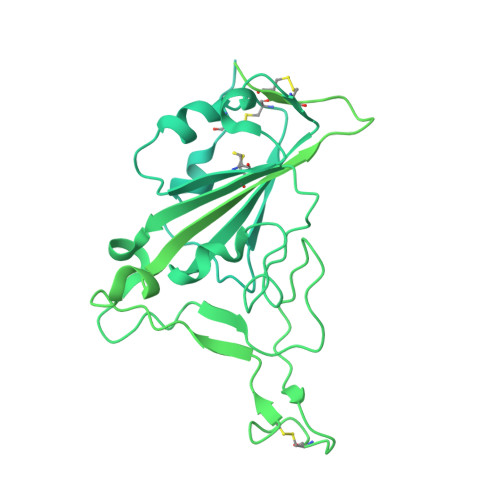
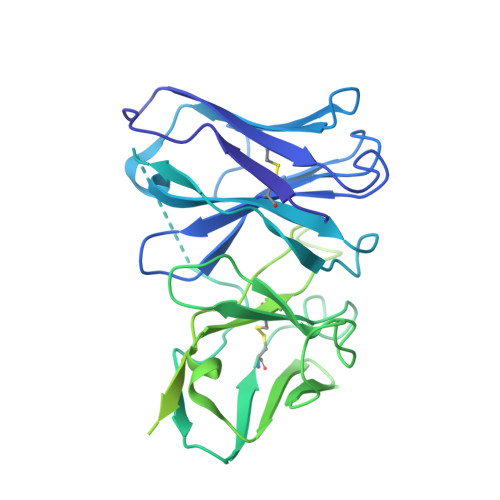
Rapid emergence of new variants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has prompted an urgent need for the development of broadly applicable and potently neutralizing antibody platform against the SARS-CoV-2, which can be used for combatting the coronavirus disease 2019 (COVID-19). In this study, based on a noncompeting pair of phage display-derived human monoclonal antibodies (mAbs) specific to the receptor-binding domain (RBD) of SARS-CoV-2 isolated from human synthetic antibody library, we generated K202.B, a novel engineered bispecific antibody with an immunoglobulin G4-single-chain variable fragment design, with sub- or low nanomolar antigen-binding avidity. Compared with the parental mAbs or mAb cocktail, the K202.B antibody showed superior neutralizing potential against a variety of SARS-CoV-2 variants in vitro. Furthermore, structural analysis of bispecific antibody-antigen complexes using cryo-electron microscopy revealed the mode of action of K202.B complexed with a fully open three-RBD-up conformation of SARS-CoV-2 trimeric spike proteins by simultaneously interconnecting two independent epitopes of the SARS-CoV-2 RBD via inter-protomer interactions. Intravenous monotherapy using K202.B exhibited potent neutralizing activity in SARS-CoV-2 wild-type- and B.1.617.2 variant-infected mouse models, without significant toxicity in vivo. The results indicate that this novel approach of development of immunoglobulin G4-based bispecific antibody from an established human recombinant antibody library is likely to be an effective strategy for the rapid development of bispecific antibodies, and timely management against fast-evolving SARS-CoV-2 variants.
- Department of Biochemistry, Kookmin University, Seoul, 02707, Republic of Korea.
Organizational Affiliation: