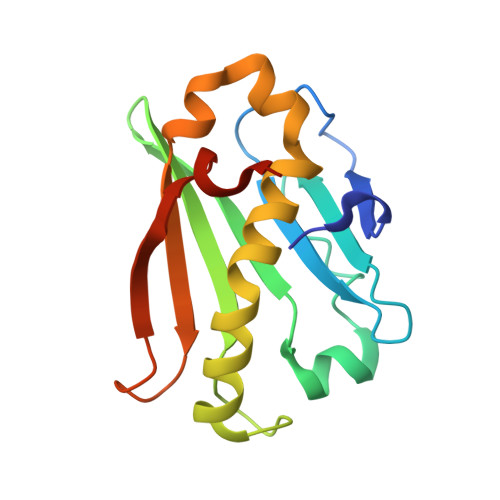
Crystal structure and reaction mechanism of a bacterial Mg-dechelatase homolog from the Chloroflexi Anaerolineae.
Dey, D., Nishijima, M., Tanaka, R., Kurisu, G., Tanaka, H., Ito, H.(2022) Protein Sci 31: e4430-e4430
- PubMed: 36173179 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4430
- Primary Citation Related Structures:
7Y5Y - PubMed Abstract:
Chlorophyll degradation plays a myriad of physiological roles in photosynthetic organisms, including acclimation to light environment and nutrient remobilization during senescence. Mg extraction from chlorophyll a is the first and committed step of the chlorophyll degradation pathway. This reaction is catalyzed by the Mg-dechelatase enzyme encoded by Stay-Green (SGR). The reaction mechanism of SGR protein remains elusive since metal ion extraction from organic molecules is not a common enzymatic reaction. Additionally, experimentally derived structural information about SGR or its homologs has not yet been reported. In this study, the crystal structure of the SGR homolog from Anaerolineae bacterium was determined using the molecular replacement method at 1.85 Å resolution. Our previous study showed that three residues-H32, D34, and D62 are essential for the catalytic activity of the enzyme. Biochemical analysis involving mutants of D34 residue further strengthened its importance in the functioning of the dechelatase. Docking simulation also revealed the interaction between the D34 side chain and central Mg ion of chlorophyll a. Structural analysis showed the arrangement of D34/H32/D62 in the form of a catalytic triad that is generally found in hydrolases. The probable reaction mechanism suggests that deprotonated D34 side chain coordinates and destabilizes Mg, resulting in Mg extraction. Besides, H32 possibly acts as a general base catalyst and D62 facilitates H32 to be a better proton acceptor. Taken together, the reaction mechanism of SGR partially mirrors the one observed in hydrolases.
- Graduate School of Life Science, Hokkaido University, Sapporo, Japan.
Organizational Affiliation: