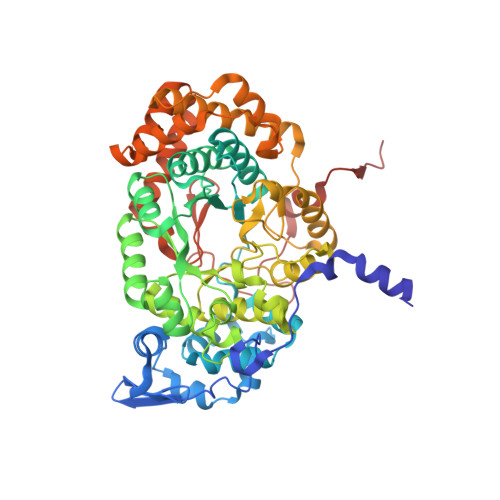
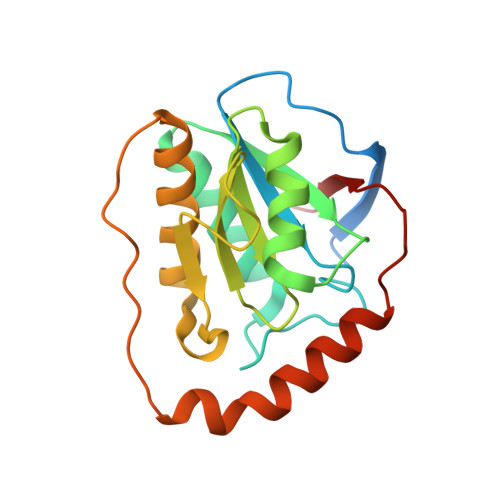
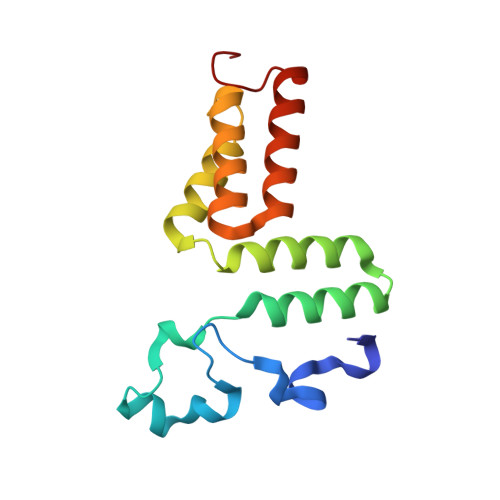
Structural Insights into the Very Low Activity of the Homocoenzyme B 12 Adenosylmethylcobalamin in Coenzyme B 12 -Dependent Diol Dehydratase and Ethanolamine Ammonia-Lyase.
Shibata, N., Higuchi, Y., Krautler, B., Toraya, T.(2022) Chemistry 28: e202202196-e202202196
- PubMed: 35974426
- DOI: https://doi.org/10.1002/chem.202202196
- Primary Citation Related Structures:
7XRK, 7XRL, 7XRM, 7XRN - PubMed Abstract:
The X-ray structures of coenzyme B 12 (AdoCbl)-dependent eliminating isomerases complexed with adenosylmethylcobalamin (AdoMeCbl) have been determined. As judged from geometries, the Co-C bond in diol dehydratase (DD) is not activated even in the presence of substrate. In ethanolamine ammonia-lyase (EAL), the bond is elongated in the absence of substrate; in the presence of substrate, the complex likely exists in both pre- and post-homolysis states. The impacts of incorporating an extra CH 2 group are different in the two enzymes: the DD active site is flexible, and AdoMeCbl binding causes large conformational changes that make DD unable to adopt the catalytic state, whereas the EAL active site is rigid, and AdoMeCbl binding does not induce significant conformational changes. Such flexibility and rigidity of the active sites might reflect the tightness of adenine binding. The structures provide good insights into the basis of the very low activity of AdoMeCbl in these enzymes.
- Department of Life Science, Graduate School of Science, University of Hyogo, 3-2-1 Koto, Kamigori-cho, Ako-gun, Hyogo, 678-1297, Japan.
Organizational Affiliation: