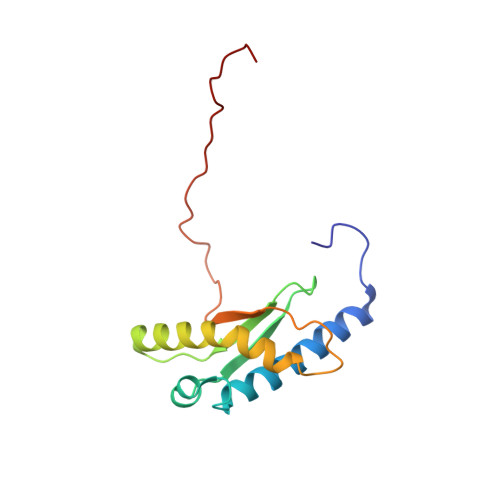
1 H, 13 C, and 15 N resonance assignments and solution structures of the KH domain of human ribosome binding factor A, mtRbfA, involved in mitochondrial ribosome biogenesis.
Kuwasako, K., Suzuki, S., Nameki, N., Takizawa, M., Takahashi, M., Tsuda, K., Nagata, T., Watanabe, S., Tanaka, A., Kobayashi, N., Kigawa, T., Guntert, P., Shirouzu, M., Yokoyama, S., Muto, Y.(2022) Biomol NMR Assign 16: 297-303
- PubMed: 35666428
- DOI: https://doi.org/10.1007/s12104-022-10094-3
- Primary Citation of Related Structures:
7X9U - PubMed Abstract:
Ribosome biogenesis is a complicated, multistage process coordinated by ribosome assembly factors. Ribosome binding factor A (RbfA) is a bacterial one, which possesses a single structural type-II KH domain. By this domain, RbfA binds to a 16S rRNA precursor in small ribosomal subunits to promote its 5'-end processing. The human RbfA homolog, mtRbfA, binds to 12S rRNAs in the mitoribosomal small subunits and promotes its critical maturation process, the dimethylation of two highly conserved consecutive adenines, which differs from that of RbfA. However, the structural basis of the mtRbfA-mediated maturation process is poorly understood. Herein, we report the 1 H, 15 N, and 13 C resonance assignments of the KH domain of mtRbfA and its solution structure. The mtRbfA domain adopts essentially the same α1-β1-β2-α2(kinked)-β3 topology as the type-II KH domain. Comparison with the RbfA counterpart showed structural differences in specific regions that function as a putative RNA-binding site. Particularly, the α2 helix of mtRbfA forms a single helix with a moderate kink at the Ser-Ala-Ala sequence, whereas the corresponding α2 helix of RbfA is interrupted by a distinct kink at the Ala-x-Gly sequence, characteristic of bacterial RbfA proteins, to adopt an α2-kink-α3 conformation. Additionally, the region linking α1 and β1 differs considerably in the sequence and structure between RbfA and mtRbfA. These findings suggest some variations of the RNA-binding mode between them and provide a structural basis for mtRbfA function in mitoribosome biogenesis.
- RIKEN Center for Life Science and Technologies, 1-7-22 Suehiro-cho, Tsurumi-ku, Yokohama, 230-0045, Japan.
Organizational Affiliation: