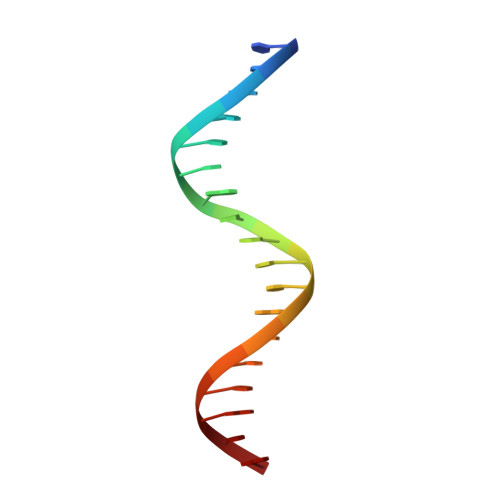
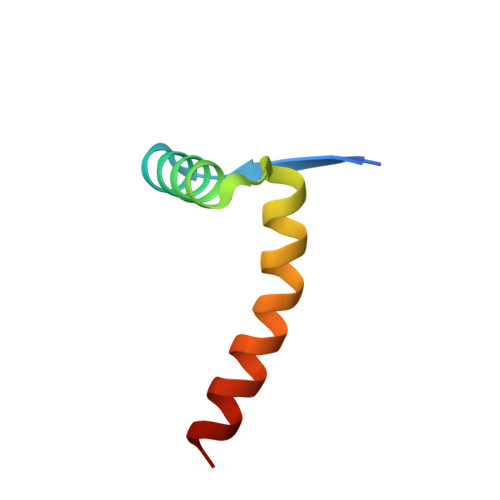
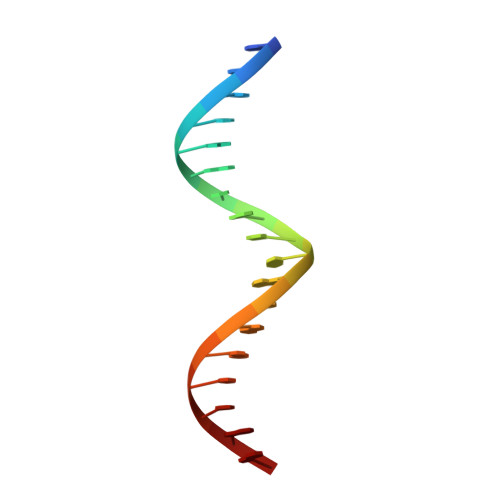
Structural and mutational analysis of MazE6-operator DNA complex provide insights into autoregulation of toxin-antitoxin systems.
Kumari, K., Sarma, S.P.(2022) Commun Biol 5: 963-963
- PubMed: 36109664 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-022-03933-5
- Primary Citation Related Structures:
7WJ0, 7WNR - PubMed Abstract:
Of the 10 paralogs of MazEF Toxin-Antitoxin system in Mycobacterium tuberculosis, MazEF6 plays an important role in multidrug tolerance, virulence, stress adaptation and Non Replicative Persistant (NRP) state establishment. The solution structures of the DNA binding domain of MazE6 and of its complex with the cognate operator DNA show that transcriptional regulation occurs by binding of MazE6 to an 18 bp operator sequence bearing the TANNNT motif (-10 region). Kinetics and thermodynamics of association, as determined by NMR and ITC, indicate that the nMazE6-DNA complex is of high affinity. Residues in N-terminal region of MazE6 that are key for its homodimerization, DNA binding specificity, and the base pairs in the operator DNA essential for the protein-DNA interaction, have been identified. It provides a basis for design of chemotherapeutic agents that will act via disruption of TA autoregulation, leading to cell death.
- Molecular Biophysics Unit, Indian Institute of Science, C. V. Raman Road, Bangalore, Karnataka, 560012, India.
Organizational Affiliation: