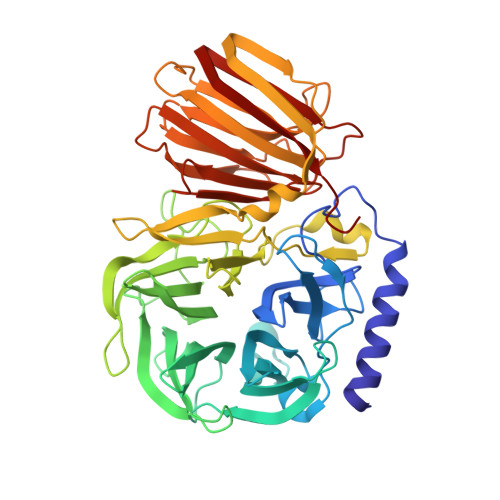
Enzymatic and structural characterization of beta-fructofuranosidase from the honeybee gut bacterium Frischella perrara.
Kubota, A., Kawai, R., Li, D., Kozono, T., Sasaki, N., Nishikawa, A., Fujii, T., Tochio, T., Tonozuka, T.(2022) Appl Microbiol Biotechnol 106: 2455-2470
- PubMed: 35267055 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00253-022-11863-9
- Primary Citation Related Structures:
7VCO, 7VCP - PubMed Abstract:
Fructooligosaccharide is a mixture of mostly the trisaccharide 1-kestose (GF 2 ), tetrasaccharide nystose (GF 3 ), and fructosyl nystose (GF 4 ). Enzymes that hydrolyze GF 3 may be useful for preparing GF 2 from the fructooligosaccharide mixture. A β-fructofuranosidase belonging to glycoside hydrolase family 32 (GH32) from the honeybee gut bacterium Frischella perrara (FperFFase) was expressed in Escherichia coli and purified. The time course of the hydrolysis of 60 mM sucrose, GF 2 , and GF 3 by FperFFase was analyzed, showing that the hydrolytic activity of FperFFase for trisaccharide GF 2 was lower than those for disaccharide sucrose and tetrasaccharide GF 3 . The crystal structure of FperFFase and its structure in complex with fructose were determined. FperFFase was found to be structurally homologous to bifidobacterial β-fructofuranosidases even though bifidobacterial enzymes preferably hydrolyze GF 2 and the amino acid residues interacting with fructose at subsite - 1 are mostly conserved between them. A proline residue was inserted between Asp298 and Ser299 using site-directed mutagenesis, and the activity of the variant 298P299 was measured. The ratio of activities for 60 mM GF 2 /GF 3 by wild-type FperFFase was 35.5%, while that of 298P299 was 23.6%, indicating that the structure of the loop comprising Trp297-Asp298-Ser299 correlated with the substrate preference of FperFFase. The crystal structure also shows that a loop consisting of residues 117-127 is likely to contribute to the substrate binding of FperFFase. The results obtained herein suggest that FperFFase is potentially useful for the manufacture of GF 2 . KEY POINTS: • Frischella β-fructofuranosidase hydrolyzed nystose more efficiently than 1-kestose. • Trp297-Asp298-Ser299 was shown to be correlated with the substrate preference. • Loop consisting of residues 117-127 appears to contribute to the substrate binding.
- Department of Applied Biological Science, Tokyo University of Agriculture and Technology, 3-5-8 Saiwai-cho, Fuchu, Tokyo, 183-8509, Japan.
Organizational Affiliation: