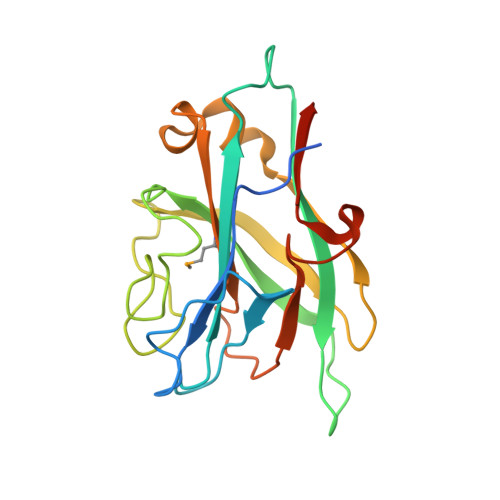
Identification and structural analysis of a carbohydrate-binding module specific to alginate, a representative of a new family, CBM96.
Ji, S., Tian, X., Li, X., She, Q.(2023) J Biological Chem 299: 102854-102854
- PubMed: 36592931
- DOI: https://doi.org/10.1016/j.jbc.2022.102854
- Primary Citation Related Structures:
7VBO - PubMed Abstract:
Carbohydrate-binding modules (CBMs) are the noncatalytic modules that assist functions of the catalytic modules in carbohydrate-active enzymes, and they are usually discrete structural domains in larger multimodular enzymes. CBMs often occur in tandem in different alginate lyases belonging to the CBM families 13, 16, and 32. However, none of the currently known CBMs in alginate lyases specifically bind to an internal alginate chain. In our investigation of the multidomain alginate lyase Dp0100 carrying several ancillary domains, we identified an alginate-binding domain denoted TM6-N4 using protein truncation analysis. The structure of this CBM domain was determined at 1.35 Å resolution. TM6-N4 exhibited an overall β-sandwich fold architecture with two antiparallel β-sheets. We identified an extended binding groove in the CBM using site-directed mutagenesis, docking, and surface electrostatic potential analysis. Affinity analysis revealed that residues of Lys10, Lys22, Lys25, Lys27, Lys31, Arg36, and Tyr159 located on the bottom or the wall of the shallow groove are responsible for alginate binding, and isothermal titration calorimetry analyses indicated that the binding cleft consists of six subsites for sugar recognition. This substrate binding pattern is typical for type B CBM, and it represents the first CBM domain that specifically binds internal alginate chain. Phylogenetic analysis supports that TM6-N4 constitutes the founding member of a new CBM family denoted as CBM96. Our reported structure not only facilitates the investigation of the CBM-alginate ligand recognition mechanism but also inspires the utilization of the CBM domain in biotechnical applications.
- CRISPR and Archaea Biology Research Center, Microbial Technology Institute and State Key Laboratory of Microbial Technology, Shandong University, Qingdao, Shandong, China. Electronic address: jisq@sdu.edu.cn.
Organizational Affiliation: