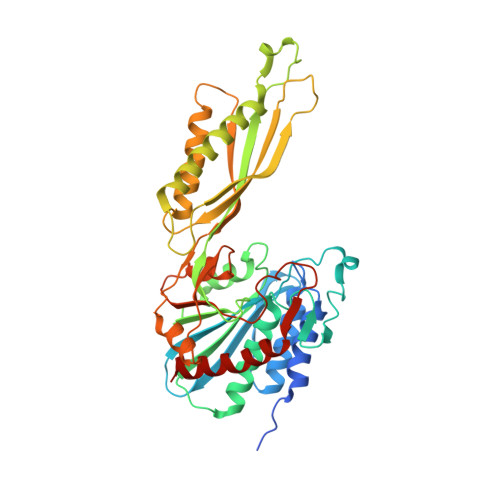
The three-dimensional structure of DapE from Enterococcus faecium reveals new insights into DapE/ArgE subfamily ligand specificity.
Terrazas-Lopez, M., Gonzalez-Segura, L., Diaz-Vilchis, A., Aguirre-Mendez, K.A., Lobo-Galo, N., Martinez-Martinez, A., Diaz-Sanchez, A.G.(2024) Int J Biol Macromol 270: 132281-132281
- PubMed: 38740150 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2024.132281
- Primary Citation Related Structures:
7UOI, 8VKT - PubMed Abstract:
DapE is a Zn 2+ -metallohydrolase recognized as a drug target for bacterial control. It is a homodimer that requires the exchange of interface strands by an induced fit essential for catalysis. Identifying novel anti-DapE agents requires greater structural details. Most of the characterized DapEs are from the Gram-negative group. Here, two high-resolution DapE crystal structures from Enterococcus faecium are presented for the first time with novel aspects. A loosened enzyme intermediate between the open and closed conformations is observed. Substrates may bind to loose state, subsequently it closes, where hydrolysis occurs, and finally, the change to the open state leads to the release of the products. Mutation of His352 suggests a role, along with His194, in the oxyanion stabilization in the mono-metalated Zn 2+ isoform, while in the di-metalated isoform, the metal center 2 complements it function. An aromatic-π box potentially involved in the interaction of DapE with other proteins, and a peptide flip could determine the specificity in the Gram-positive ArgE/DapE group. Finally, details of two extra-catalytic cavities whose geometry changes depending on the conformational state of the enzyme are presented. These cavities could be a target for developing non-competitive agents that trap the enzyme in an inactive state.
- Universidad Autónoma de Ciudad Juárez, Ciudad Juárez, Instituto de Ciencias Biomédicas, Departamento de Ciencias Químico-Biológicas, Chihuahua, CP 32310, Mexico.
Organizational Affiliation: