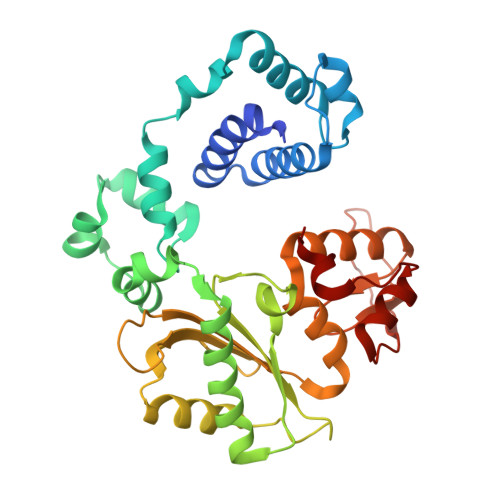
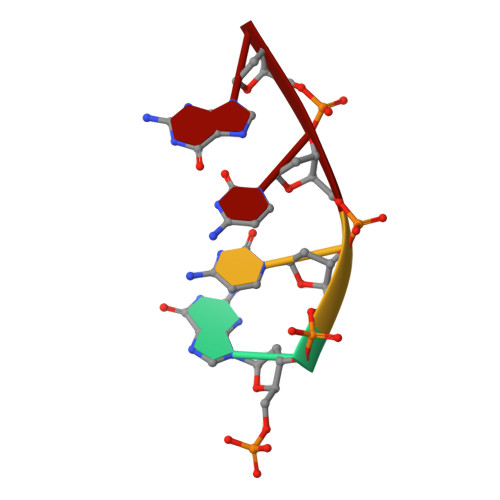
Pol lambda promotes microhomology-mediated end-joining.
Chandramouly, G., Jamsen, J., Borisonnik, N., Tyagi, M., Calbert, M.L., Tredinnick, T., Ozdemir, A.Y., Kent, T., Demidova, E.V., Arora, S., Wilson, S.H., Pomerantz, R.T.(2023) Nat Struct Mol Biol 30: 107-114
- PubMed: 36536104 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-022-00895-4
- Primary Citation Related Structures:
7UN7 - PubMed Abstract:
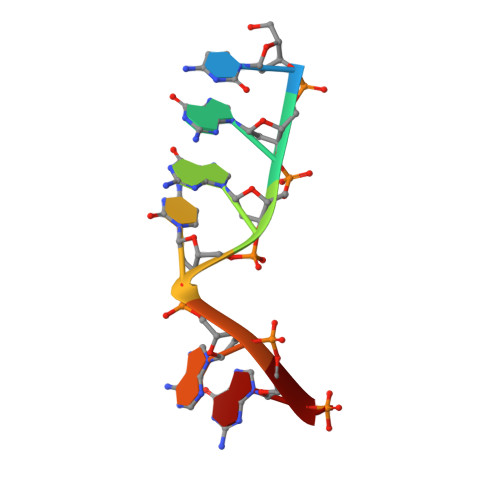
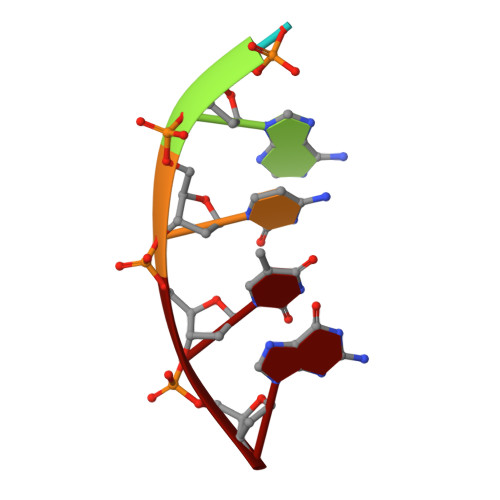
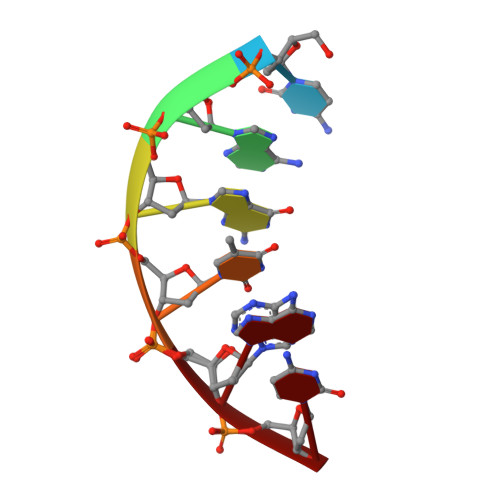
The double-strand break (DSB) repair pathway called microhomology-mediated end-joining (MMEJ) is thought to be dependent on DNA polymerase theta (Polθ) and occur independently of nonhomologous end-joining (NHEJ) factors. An unresolved question is whether MMEJ is facilitated by a single Polθ-mediated end-joining pathway or consists of additional undiscovered pathways. We find that human X-family Polλ, which functions in NHEJ, additionally exhibits robust MMEJ activity like Polθ. Polλ promotes MMEJ in mammalian cells independently of essential NHEJ factors LIG4/XRCC4 and Polθ, which reveals a distinct Polλ-dependent MMEJ mechanism. X-ray crystallography employing in situ photo-induced DSB formation captured Polλ in the act of stabilizing a microhomology-mediated DNA synapse with incoming nucleotide at 2.0 Å resolution and reveals how Polλ performs replication across a DNA synapse joined by minimal base-pairing. Last, we find that Polλ is semisynthetic lethal with BRCA1 and BRCA2. Together, these studies indicate Polλ MMEJ as a distinct DSB repair mechanism.
- Department of Biochemistry and Molecular Biology, Sidney Kimmel Cancer Center, Thomas Jefferson University, Philadelphia, PA, USA.
Organizational Affiliation: