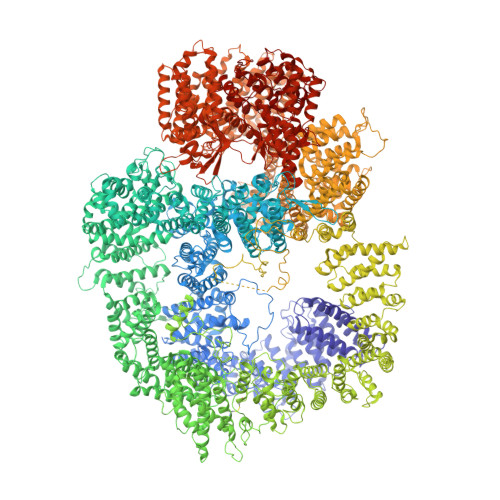
Structural analysis of the basal state of the Artemis:DNA-PKcs complex.
Watanabe, G., Lieber, M.R., Williams, D.R.(2022) Nucleic Acids Res 50: 7697-7720
- PubMed: 35801871 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac564
- Primary Citation Related Structures:
7TYR - PubMed Abstract:
Artemis nuclease and DNA-dependent protein kinase catalytic subunit (DNA-PKcs) are key components in nonhomologous DNA end joining (NHEJ), the major repair mechanism for double-strand DNA breaks. Artemis activation by DNA-PKcs resolves hairpin DNA ends formed during V(D)J recombination. Artemis deficiency disrupts development of adaptive immunity and leads to radiosensitive T- B- severe combined immunodeficiency (RS-SCID). An activated state of Artemis in complex with DNA-PK was solved by cryo-EM recently, which showed Artemis bound to the DNA. Here, we report that the pre-activated form (basal state) of the Artemis:DNA-PKcs complex is stable on an agarose-acrylamide gel system, and suitable for cryo-EM structural analysis. Structures show that the Artemis catalytic domain is dynamically positioned externally to DNA-PKcs prior to ABCDE autophosphorylation and show how both the catalytic and regulatory domains of Artemis interact with the N-HEAT and FAT domains of DNA-PKcs. We define a mutually exclusive binding site for Artemis and XRCC4 on DNA-PKcs and show that an XRCC4 peptide disrupts the Artemis:DNA-PKcs complex. All of the findings are useful in explaining how a hypomorphic L3062R missense mutation of DNA-PKcs could lead to insufficient Artemis activation, hence RS-SCID. Our results provide various target site candidates to design disruptors for Artemis:DNA-PKcs complex formation.
- Department of Pathology, Department of Biochemistry & Molecular Biology, Department of Molecular Microbiology & Immunology, and Section of Computational & Molecular Biology, USC Norris Comprehensive Cancer Center, University of Southern California Keck School of Medicine, 1441 Eastlake Ave, Rm. 5428, Los Angeles, CA 90089, USA.
Organizational Affiliation: