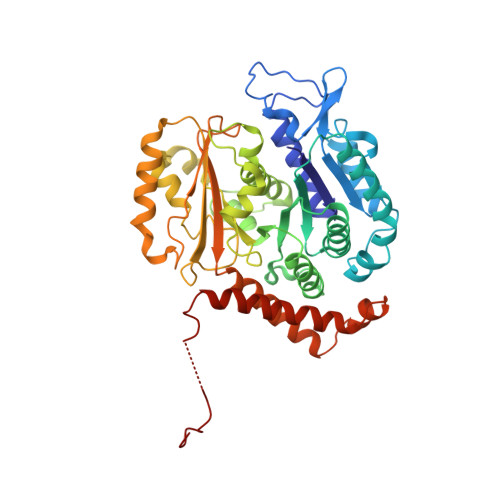
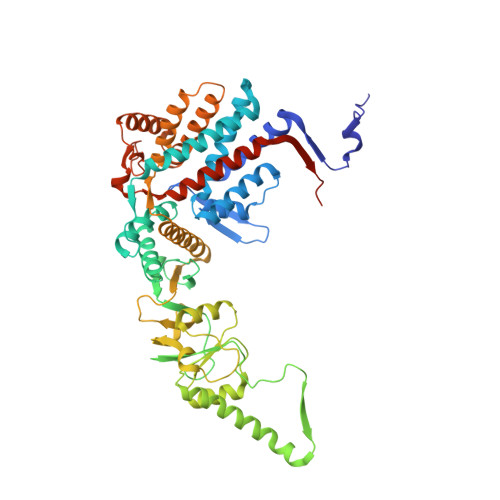
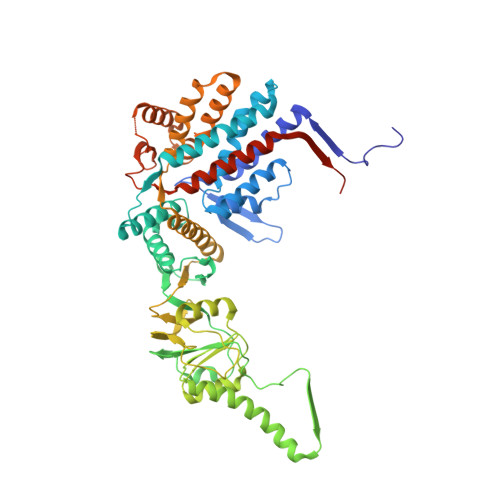
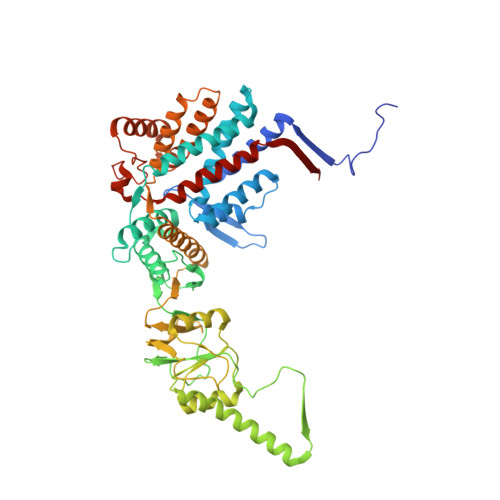
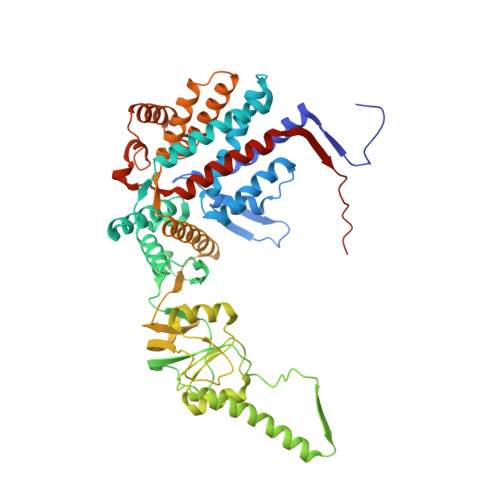
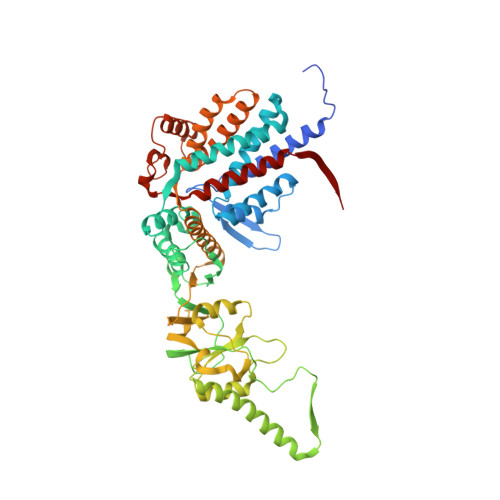
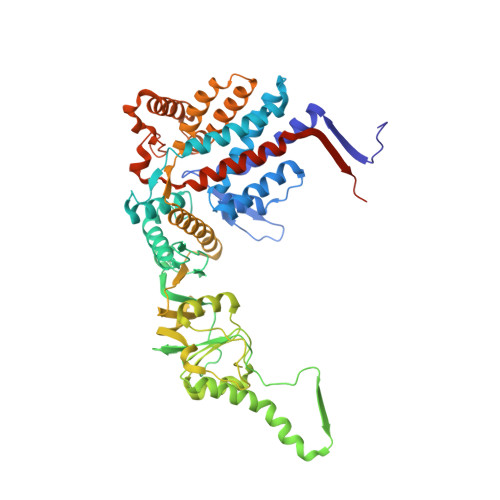
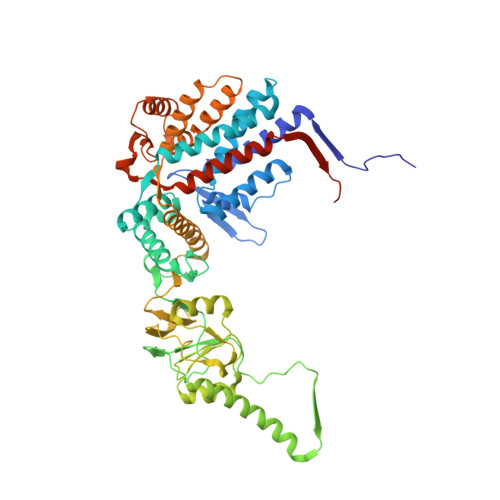
Structural visualization of the tubulin folding pathway directed by human chaperonin TRiC/CCT.
Gestaut, D., Zhao, Y., Park, J., Ma, B., Leitner, A., Collier, M., Pintilie, G., Roh, S.H., Chiu, W., Frydman, J.(2022) Cell 185: 4770-4787.e20
- PubMed: 36493755
- DOI: https://doi.org/10.1016/j.cell.2022.11.014
- Primary Citation Related Structures:
7TRG, 7TTN, 7TTT, 7TUB, 7WU7 - PubMed Abstract:
The ATP-dependent ring-shaped chaperonin TRiC/CCT is essential for cellular proteostasis. To uncover why some eukaryotic proteins can only fold with TRiC assistance, we reconstituted the folding of β-tubulin using human prefoldin and TRiC. We find unstructured β-tubulin is delivered by prefoldin to the open TRiC chamber followed by ATP-dependent chamber closure. Cryo-EM resolves four near-atomic-resolution structures containing progressively folded β-tubulin intermediates within the closed TRiC chamber, culminating in native tubulin. This substrate folding pathway appears closely guided by site-specific interactions with conserved regions in the TRiC chamber. Initial electrostatic interactions between the TRiC interior wall and both the folded tubulin N domain and its C-terminal E-hook tail establish the native substrate topology, thus enabling C-domain folding. Intrinsically disordered CCT C termini within the chamber promote subsequent folding of tubulin's core and middle domains and GTP-binding. Thus, TRiC's chamber provides chemical and topological directives that shape the folding landscape of its obligate substrates.
- Department of Biology, Stanford University, Stanford, CA 94305, USA.
Organizational Affiliation: