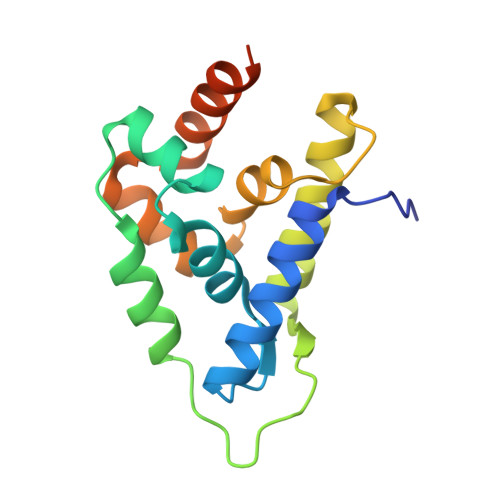
Atomic structure of the Leishmania spp. Hsp100 N-domain.
Mercado, J.M., Lee, S., Chang, C., Sung, N., Soong, L., Catic, A., Tsai, F.T.F.(2022) Proteins 90: 1242-1246
- PubMed: 35122310 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.26310
- Primary Citation Related Structures:
7TFM - PubMed Abstract:
Hsp100 is an ATP-dependent unfoldase that promotes protein disaggregation or facilitates the unfolding of aggregation-prone polypeptides marked for degradation. Recently, new Hsp100 functions are emerging. In Plasmodium, an Hsp100 drives malaria protein export, presenting a novel drug target. Whether Hsp100 has a similar function in other protists is unknown. We present the 1.06 Å resolution crystal structure of the Hsp100 N-domain from Leishmania spp., the causative agent of leishmaniasis in humans. Our structure reveals a network of methionines and aromatic amino acids that define the putative substrate-binding site and likely evolved to protect Hsp100 from oxidative damage in host immune cells.
- Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Organizational Affiliation: