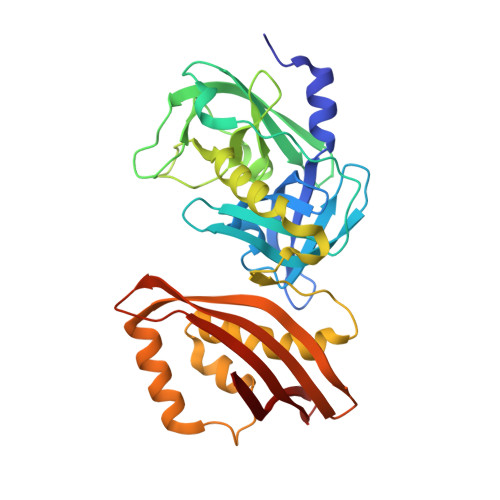
The structure of the Clostridium thermocellum RsgI9 ectodomain provides insight into the mechanism of biomass sensing.
Mahoney, B.J., Takayesu, A., Zhou, A., Cascio, D., Clubb, R.T.(2022) Proteins 90: 1457-1467
- PubMed: 35194841
- DOI: https://doi.org/10.1002/prot.26326
- Primary Citation Related Structures:
7SJY - PubMed Abstract:
Clostridium thermocellum is actively being developed as a microbial platform to produce biofuels and chemicals from renewable plant biomass. An attractive feature of this bacterium is its ability to efficiently degrade lignocellulose using surface-displayed cellulosomes, large multi-protein complexes that house different types of cellulase enzymes. Clostridium thermocellum tailors the enzyme composition of its cellulosome using nine membrane-embedded anti-σ factors (RsgI1-9), which are thought to sense different types of extracellular carbohydrates and then elicit distinct gene expression programs via cytoplasmic σ factors. Here we show that the RsgI9 anti-σ factor interacts with cellulose via a C-terminal bi-domain unit. A 2.0 Å crystal structure reveals that the unit is constructed from S1C peptidase and NTF2-like protein domains that contain a potential binding site for cellulose. Small-angle X-ray scattering experiments of the intact ectodomain indicate that it adopts a bi-lobed, elongated conformation. In the structure, a conserved RsgI extracellular (CRE) domain is connected to the bi-domain via a proline-rich linker, which is expected to project the carbohydrate-binding unit ~160 Å from the cell surface. The CRE and proline-rich elements are conserved in several other C. thermocellum anti-σ factors, suggesting that they will also form extended structures that sense carbohydrates.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, Los Angeles, California, USA.
Organizational Affiliation: