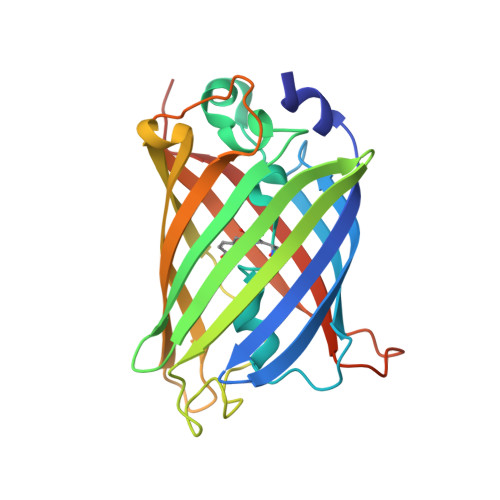
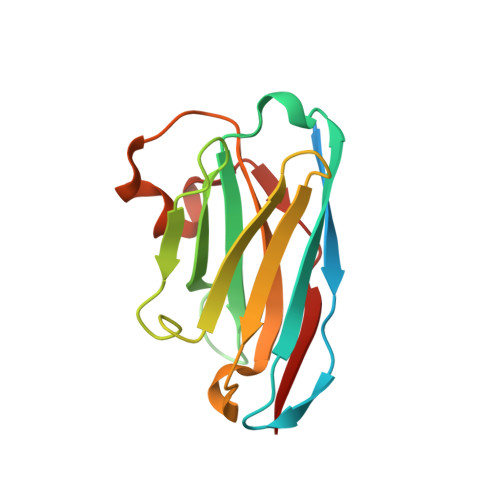
High-efficiency recombinant protein purification using mCherry and YFP nanobody affinity matrices.
Cong, A.T.Q., Witter, T.L., Schellenberg, M.J.(2022) Protein Sci 31: e4383-e4383
- PubMed: 36040252
- DOI: https://doi.org/10.1002/pro.4383
- Primary Citation Related Structures:
7SAH, 7SAI, 7SAJ, 7SAK, 7SAL - PubMed Abstract:
Mammalian cell lines are important expression systems for large proteins and protein complexes, particularly when the acquisition of post-translational modifications in the protein's native environment is desired. However, low or variable transfection efficiencies are challenges that must be overcome to use such an expression system. Expression of recombinant proteins as a fluorescent protein fusion enables real-time monitoring of protein expression, and also provides an affinity handle for one-step protein purification using a suitable affinity reagent. Here, we describe a panel of anti-GFP and anti-mCherry nanobody affinity matrices and their efficacy for purification of GFP/YFP or mCherry fusion proteins. We define the molecular basis by which they bind their target proteins using X-ray crystallography. From these analyses, we define an optimal pair of nanobodies for purification of recombinant protein tagged with GFP/YFP or mCherry, and demonstrate these nanobody-sepharose supports are stable to many rounds of cleaning and extended incubation in denaturing conditions. Finally, we demonstrate the utility of the mCherry-tag system by using it to purify recombinant human topoisomerase 2α expressed in HEK293F cells. The mCherry-tag and GFP/YFP-tag expression systems can be utilized for recombinant protein expression individually or in tandem for mammalian protein expression systems where real-time monitoring of protein expression levels and a high-efficiency purification step is needed.
- Department of Biochemistry and Molecular Biology, Mayo Clinic, Rochester, Minnesota, USA.
Organizational Affiliation: