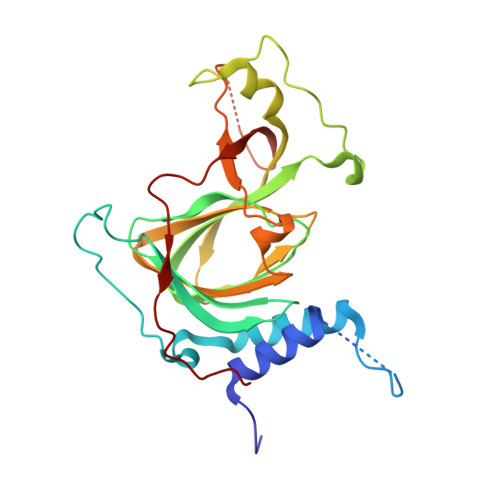
Crystal structure of human cysteamine dioxygenase provides a structural rationale for its function as an oxygen sensor.
Wang, Y., Shin, I., Li, J., Liu, A.(2021) J Biological Chem 297: 101176-101176
- PubMed: 34508780
- DOI: https://doi.org/10.1016/j.jbc.2021.101176
- Primary Citation Related Structures:
7REI - PubMed Abstract:
Cysteamine dioxygenase (ADO) plays a vital role in regulating thiol metabolism and preserving oxygen homeostasis in humans by oxidizing the sulfur of cysteamine and N-terminal cysteine-containing proteins to their corresponding sulfinic acids using O 2 as a cosubstrate. However, as the only thiol dioxygenase that processes both small-molecule and protein substrates, how ADO handles diverse substrates of disparate sizes to achieve various reactions is not understood. The knowledge gap is mainly due to the three-dimensional structure not being solved, as ADO cannot be directly compared with other known thiol dioxygenases. Herein, we report the first crystal structure of human ADO at a resolution of 1.78 Å with a nickel-bound metal center. Crystallization was achieved through both metal substitution and C18S/C239S double mutations. The metal center resides in a tunnel close to an entry site flanked by loops. While ADO appears to use extensive flexibility to handle substrates of different sizes, it also employs proline and proline pairs to maintain the core protein structure and to retain the residues critical for catalysis in place. This feature distinguishes ADO from thiol dioxygenases that only oxidize small-molecule substrates, possibly explaining its divergent substrate specificity. Our findings also elucidate the structural basis for ADO functioning as an oxygen sensor by modifying N-degron substrates to transduce responses to hypoxia. Thus, this work fills a gap in structure-function relationships of the thiol dioxygenase family and provides a platform for further mechanistic investigation and therapeutic intervention targeting impaired oxygen sensing.
- Department of Chemistry, The University of Texas at San Antonio, Texas, USA.
Organizational Affiliation: