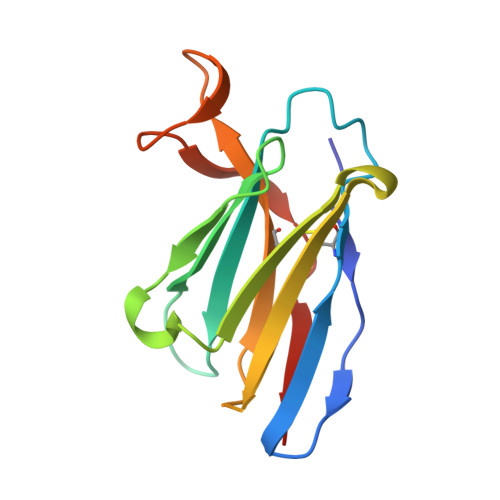
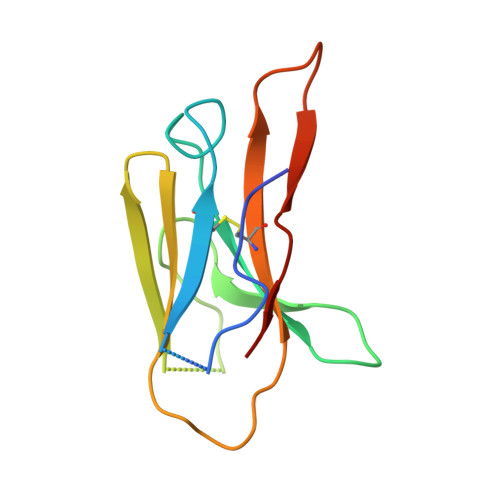
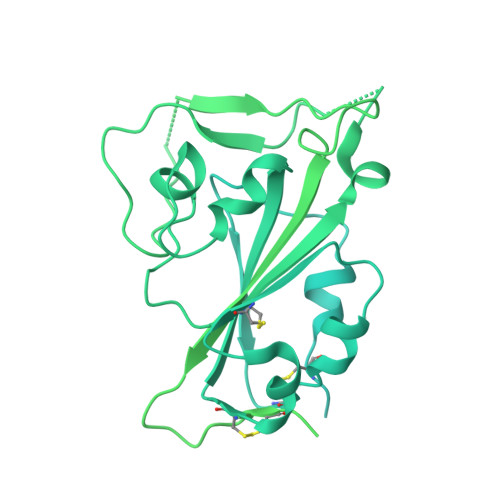
Broad sarbecovirus neutralization by a human monoclonal antibody.
Tortorici, M.A., Czudnochowski, N., Starr, T.N., Marzi, R., Walls, A.C., Zatta, F., Bowen, J.E., Jaconi, S., Di Iulio, J., Wang, Z., De Marco, A., Zepeda, S.K., Pinto, D., Liu, Z., Beltramello, M., Bartha, I., Housley, M.P., Lempp, F.A., Rosen, L.E., Dellota Jr., E., Kaiser, H., Montiel-Ruiz, M., Zhou, J., Addetia, A., Guarino, B., Culap, K., Sprugasci, N., Saliba, C., Vetti, E., Giacchetto-Sasselli, I., Fregni, C.S., Abdelnabi, R., Foo, S.C., Havenar-Daughton, C., Schmid, M.A., Benigni, F., Cameroni, E., Neyts, J., Telenti, A., Virgin, H.W., Whelan, S.P.J., Snell, G., Bloom, J.D., Corti, D., Veesler, D., Pizzuto, M.S.(2021) Nature 597: 103-108
- PubMed: 34280951 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03817-4
- Primary Citation Related Structures:
7RA8, 7RAL - PubMed Abstract:
The recent emergence of SARS-CoV-2 variants of concern 1-10 and the recurrent spillovers of coronaviruses 11,12 into the human population highlight the need for broadly neutralizing antibodies that are not affected by the ongoing antigenic drift and that can prevent or treat future zoonotic infections. Here we describe a human monoclonal antibody designated S2X259, which recognizes a highly conserved cryptic epitope of the receptor-binding domain and cross-reacts with spikes from all clades of sarbecovirus. S2X259 broadly neutralizes spike-mediated cell entry of SARS-CoV-2, including variants of concern (B.1.1.7, B.1.351, P.1, and B.1.427/B.1.429), as well as a wide spectrum of human and potentially zoonotic sarbecoviruses through inhibition of angiotensin-converting enzyme 2 (ACE2) binding to the receptor-binding domain. Furthermore, deep-mutational scanning and in vitro escape selection experiments demonstrate that S2X259 possesses an escape profile that is limited to a single substitution, G504D. We show that prophylactic and therapeutic administration of S2X259 protects Syrian hamsters (Mesocricetus auratus) against challenge with the prototypic SARS-CoV-2 and the B.1.351 variant of concern, which suggests that this monoclonal antibody is a promising candidate for the prevention and treatment of emergent variants and zoonotic infections. Our data reveal a key antigenic site that is targeted by broadly neutralizing antibodies and will guide the design of vaccines that are effective against all sarbecoviruses.
- Department of Biochemistry, University of Washington, Seattle, WA, USA.
Organizational Affiliation: