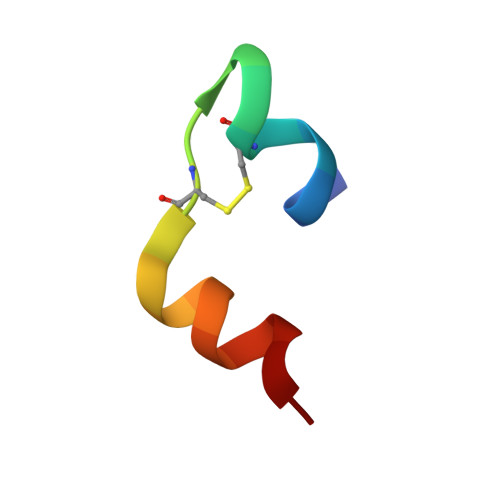
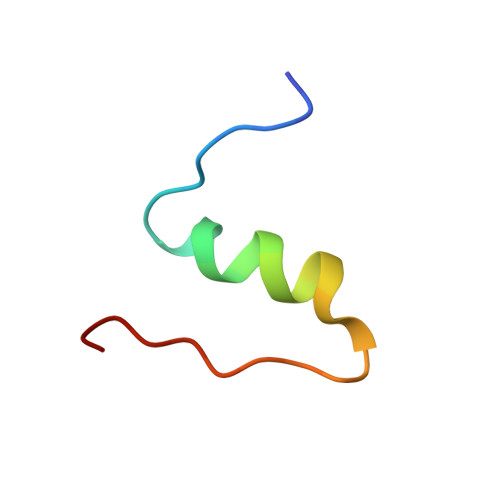
The T 2 structure of polycrystalline cubic human insulin.
Triandafillidis, D.P., Karavassili, F., Spiliopoulou, M., Valmas, A., Athanasiadou, M., Nikolaras, G., Fili, S., Kontou, P., Bowler, M.W., Chasapis, C.T., Von Dreele, R.B., Fitch, A.N., Margiolaki, I.(2023) Acta Crystallogr D Struct Biol 79: 374-386
- PubMed: 37039669 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798323001328
- Primary Citation Related Structures:
7QAC - PubMed Abstract:
The polymorphism of human insulin upon pH variation was characterized via X-ray powder diffraction, employing a crystallization protocol previously established for co-crystallization with phenolic derivatives. Two distinct rhombohedral (R3) polymorphs and one cubic (I2 1 3) polymorph were identified with increasing pH, corresponding to the T 6 , T 3 R 3 f and T 2 conformations of insulin, respectively. The structure of the cubic T 2 polymorph was determined via multi-profile stereochemically restrained Rietveld refinement at 2.7 Å resolution. This constitutes the first cubic insulin structure to be determined from crystals grown in the presence of zinc ions, although no zinc binding was observed. The differences of the polycrystalline variant from other cubic insulin structures, as well as the nature of the pH-driven phase transitions, are discussed in detail.
- Section of Genetics, Cell Biology and Development, Department of Biology, University of Patras, 26504 Patras, Greece.
Organizational Affiliation: