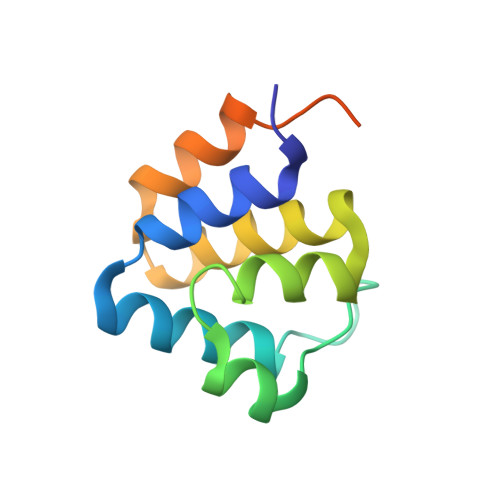
Directionality of PYD filament growth determined by the transition of NLRP3 nucleation seeds to ASC elongation.
Hochheiser, I.V., Behrmann, H., Hagelueken, G., Rodriguez-Alcazar, J.F., Kopp, A., Latz, E., Behrmann, E., Geyer, M.(2022) Sci Adv 8: eabn7583-eabn7583
- PubMed: 35559676 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abn7583
- Primary Citation Related Structures:
7PZD - PubMed Abstract:
Inflammasomes sense intrinsic and extrinsic danger signals to trigger inflammatory responses and pyroptotic cell death. Homotypic pyrin domain (PYD) interactions of inflammasome forming nucleotide-binding oligomerization domain (NOD)-like receptors with the adaptor protein ASC (apoptosis-associated speck-like protein containing a CARD) mediate oligomerization into filamentous assemblies. We describe the cryo-electron microscopy (cryo-EM) structure of the human NLRP3 PYD filament and identify a pattern of highly polar interface residues that form the homomeric interactions leading to characteristic filament ends designated as A- and B-ends. Coupling a titration polymerization assay to cryo-EM, we demonstrate that ASC adaptor protein elongation on NLRP3 PYD nucleation seeds is unidirectional, associating exclusively to the B-end of the filament. Notably, NLRP3 and ASC PYD filaments exhibit the same symmetry in rotation and axial rise per subunit, allowing a continuous transition between NLRP3 and ASC. Integrating the directionality of filament growth, we present a molecular model of the ASC speck consisting of active NLRP3, ASC, and Caspase-1 proteins.
- Institute of Structural Biology, University of Bonn, Venusberg-Campus 1, 53127 Bonn, Germany.
Organizational Affiliation: