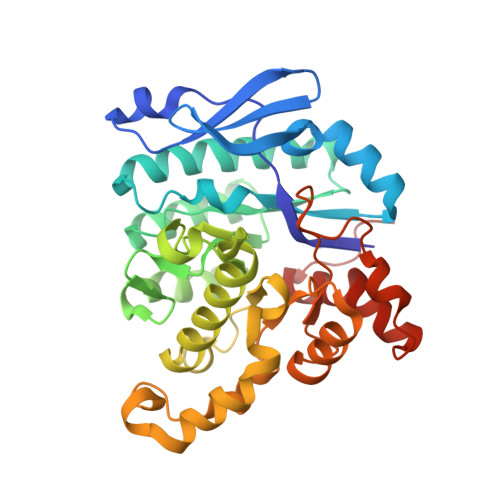
Structural Basis of Human Dimeric alpha-Amino-beta-Carboxymuconate-epsilon-Semialdehyde Decarboxylase Inhibition With TES-1025.
Cianci, M., Giacche, N., Cialabrini, L., Carotti, A., Liscio, P., Rosatelli, E., De Franco, F., Gasparrini, M., Robertson, J., Amici, A., Raffaelli, N., Pellicciari, R.(2022) Front Mol Biosci 9: 834700-834700
- PubMed: 35463964
- DOI: https://doi.org/10.3389/fmolb.2022.834700
- Primary Citation Related Structures:
7PWY - PubMed Abstract:
Human α-amino-β-carboxymuconate-ε-semialdehyde decarboxylase (ACMSD) stands at a branch point of the de novo NAD + synthesis pathway and plays an important role in maintaining NAD + homeostasis. It has been recently identified as a novel therapeutic target for a wide range of diseases, including inflammatory, metabolic disorders, and aging. So far, in absence of potent and selective enzyme inhibitors, only a crystal structure of the complex of human dimeric ACMSD with pseudo-substrate dipicolinic acid has been resolved. In this study, we report the crystal structure of the complex of human dimeric ACMSD with TES-1025, the first nanomolar inhibitor of this target, which shows a binding conformation different from the previously published predicted binding mode obtained by docking experiments. The inhibitor has a K i value of 0.85 ± 0.22 nM and binds in the catalytic site, interacting with the Zn 2+ metal ion and with residues belonging to both chains of the dimer. The results provide new structural information about the mechanism of inhibition exerted by a novel class of compounds on the ACMSD enzyme, a novel therapeutic target for liver and kidney diseases.
- Biochemistry and Structural Biology Laboratory, Department of Agricultural, Food and Environmental Sciences, Polytechnic University of Marche, Ancona, Italy.
Organizational Affiliation: