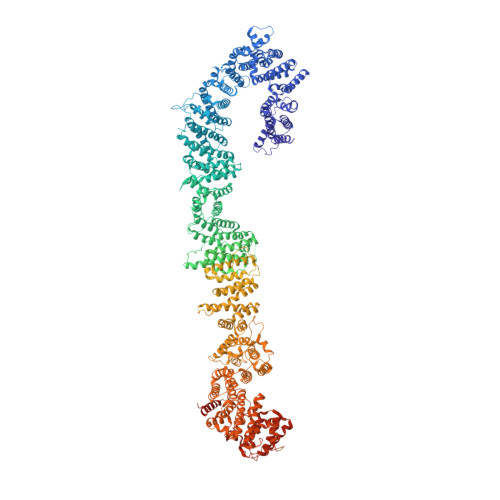
The structure of neurofibromin isoform 2 reveals different functional states.
Naschberger, A., Baradaran, R., Rupp, B., Carroni, M.(2021) Nature 599: 315-319
- PubMed: 34707296 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-04024-x
- Primary Citation Related Structures:
7PGP, 7PGQ, 7PGR, 7PGS, 7PGT, 7PGU - PubMed Abstract:
The autosomal dominant monogenetic disease neurofibromatosis type 1 (NF1) affects approximately one in 3,000 individuals and is caused by mutations in the NF1 tumour suppressor gene, leading to dysfunction in the protein neurofibromin (Nf1) 1,2 . As a GTPase-activating protein, a key function of Nf1 is repression of the Ras oncogene signalling cascade. We determined the human Nf1 dimer structure at an overall resolution of 3.3 Å. The cryo-electron microscopy structure reveals domain organization and structural details of the Nf1 exon 23a splicing 3 isoform 2 in a closed, self-inhibited, Zn-stabilized state and an open state. In the closed conformation, HEAT/ARM core domains shield the GTPase-activating protein-related domain (GRD) so that Ras binding is sterically inhibited. In a distinctly different, open conformation of one protomer, a large-scale movement of the GRD occurs, which is necessary to access Ras, whereas Sec14-PH reorients to allow interaction with the cellular membrane 4 . Zn incubation of Nf1 leads to reduced Ras-GAP activity with both protomers in the self-inhibited, closed conformation stabilized by a Zn binding site between the N-HEAT/ARM domain and the GRD-Sec14-PH linker. The transition between closed, self-inhibited states of Nf1 and open states provides guidance for targeted studies deciphering the complex molecular mechanism behind the widespread neurofibromatosis syndrome and Nf1 dysfunction in carcinogenesis.
- SciLifeLab, Department of Biochemistry and Biophysics, Stockholm University, Solna, Sweden.
Organizational Affiliation: