X-ray structure of a cystein mutant of Cyclophilin A tethered to an aromatic oligoamide foldamer complexed with Cyclosporin A
Fischer, L., Savko, M., Langlois d'Estaintot, B., Buratto, J., Vallade, M., Huc, I.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Peptidyl-prolyl cis-trans isomerase A | A, C [auth D] | 165 | Homo sapiens | Mutation(s): 1 Gene Names: PPIA, CYPA EC: 5.2.1.8 | 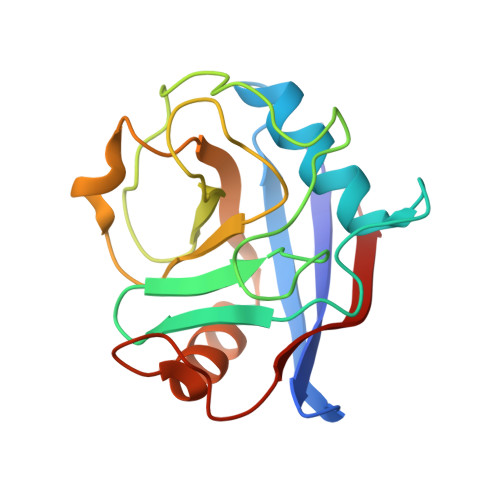 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P62937 GTEx: ENSG00000196262 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P62937 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cyclosporin A | B, D [auth E] | 11 | Tolypocladium inflatum | Mutation(s): 0 | 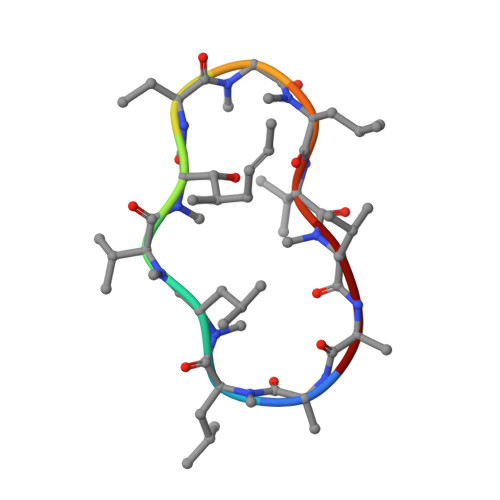 |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Aromatic foldamer | E [auth F], F [auth C] | 7 | synthetic construct | Mutation(s): 0 |  |
| Modified Residues 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| ABA Query on ABA | B, D [auth E] | L-PEPTIDE LINKING | C4 H9 N O2 |  | ALA |
| BMT Query on BMT | B, D [auth E] | L-PEPTIDE LINKING | C10 H19 N O3 |  | THR |
| DAL Query on DAL | B, D [auth E] | D-PEPTIDE LINKING | C3 H7 N O2 |  | -- |
| MLE Query on MLE | B, D [auth E] | L-PEPTIDE LINKING | C7 H15 N O2 |  | LEU |
| MVA Query on MVA | B, D [auth E] | L-PEPTIDE LINKING | C6 H13 N O2 |  | VAL |
| SAR Query on SAR | B, D [auth E] | PEPTIDE LINKING | C3 H7 N O2 |  | GLY |
| QUJ Query on QUJ | E [auth F], F [auth C] | L-PEPTIDE LINKING | C14 H16 N2 O3 |  | -- |
| QUK Query on QUK | E [auth F], F [auth C] | L-PEPTIDE LINKING | C13 H15 N3 O3 |  | -- |
| QVS Query on QVS | E [auth F], F [auth C] | L-PEPTIDE LINKING | C10 H8 N2 O3 |  | -- |
| Entity ID: 2 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_000142 Query on PRD_000142 | B, D [auth E] | Cyclosporin A | Cyclic peptide / Immunosuppressant |  | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 42.474 | α = 90 |
| b = 70.891 | β = 102.7 |
| c = 64.09 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Other government | France | -- |