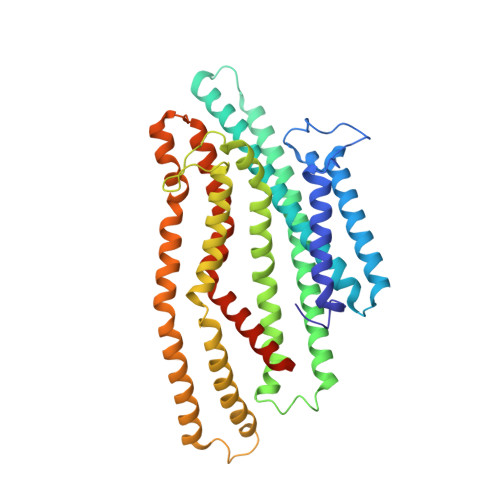
Molecular mechanism of SbmA, a promiscuous transporter exploited by antimicrobial peptides.
Ghilarov, D., Inaba-Inoue, S., Stepien, P., Qu, F., Michalczyk, E., Pakosz, Z., Nomura, N., Ogasawara, S., Walker, G.C., Rebuffat, S., Iwata, S., Heddle, J.G., Beis, K.(2021) Sci Adv 7: eabj5363-eabj5363
- PubMed: 34516884 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abj5363
- Primary Citation Related Structures:
7P34 - PubMed Abstract:
Antibiotic metabolites and antimicrobial peptides mediate competition between bacterial species. Many of them hijack inner and outer membrane proteins to enter cells. Sensitivity of enteric bacteria to multiple peptide antibiotics is controlled by the single inner membrane protein SbmA. To establish the molecular mechanism of peptide transport by SbmA and related BacA, we determined their cryo–electron microscopy structures at 3.2 and 6 Å local resolution, respectively. The structures show a previously unknown fold, defining a new class of secondary transporters named SbmA-like peptide transporters. The core domain includes conserved glutamates, which provide a pathway for proton translocation, powering transport. The structures show an outward-open conformation with a large cavity that can accommodate diverse substrates. We propose a molecular mechanism for antibacterial peptide uptake paving the way for creation of narrow-targeted therapeutics.
- Malopolska Centre of Biotechnology, Jagiellonian University, Krakow, Poland.
Organizational Affiliation: